‘Can you hear me, now?’ A new strategy ‘raises the volume’ of hard-to-detect gut-body communication
Throughout the gastrointestinal tract there are specialized hormone-producing cells called enteroendocrine cells and, although they comprise only a small population of the total cells, they are one of the most important moderators of communication between the gut and the rest of the body. Studying these cells, however, has been difficult.
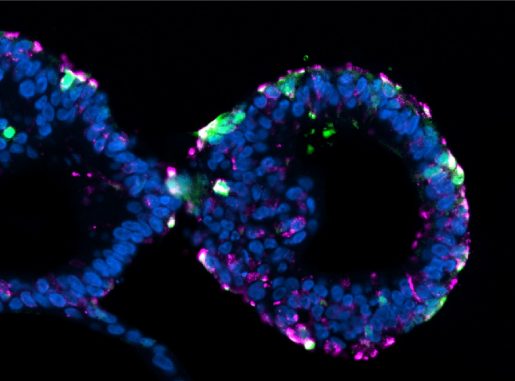
“Enteroendocrine cells are extremely challenging to study because we just don’t have a lot of cells,” said co-corresponding author Dr. Joseph M. Hyser, assistant professor of virology and microbiology and member of the Alkek Center for Metagenomics and Microbiome Research and the Dan L Duncan Comprehensive Cancer Center at Baylor College of Medicine. “They represent less than 1 percent of all the cells in the intestinal epithelium.”

“To circumvent this limitation, researchers have used cancer-derived cell lines,” said Dr. Alexandra L. Chang-Graham, co-first author of the paper, who was a student in the Medical Scientist Training Program (M.D./Ph.D.) in the Hyser lab while she was working on this project. “However, although the cancer-derived models recapitulate some of the abilities of the enteroendocrine cells, it is uncertain how much they really reproduce them.”
‘Raising the volume’
Looking to find a strategy that would enable them to study these cells, Hyser and Chang-Graham partnered up with other researchers at Baylor and Texas Children’s Hospital to develop a strategy that would increase the number of cells.
They worked with a type of human intestinal epithelial cell culture system called enteroids that is set up in Dr. Mary Estes’s lab at Baylor to grow and study norovirus. These multi-cellular cultures are made from adult intestinal stem cells from patient tissues, which naturally contain few enteroendocrine cells.
To expand the population of enteroendocrine cells, the researchers built on previous work showing that overexpressing transcription factor neurogenin-3 drives stem cells in the gut into becoming enteroendocrine cells.

“We genetically engineered human intestinal enteroids to express gene neurogenin-3 that is inducible by doxycycline,” Chang-Graham said. “Doxycycline worked as a switch. When added to the cultures, it turned on the production of neurogenin-3, which in turn triggered the expression of many other genes and the development of endocrine enteroid cells. This strategy resulted in an expansion of the endocrine enteroid cell population from one to 40 percent, while the numbers of other cell types of the epithelium remained largely unchanged.”
The expanded endocrine enteroid cell population responded to hormonal and viral stimuli as native endocrine cells in the intestine do; they secreted serotonin and other hormonal and neurotransmitter mediators, but now their levels were easily detectable.
Opening the doors to new research opportunities
“Now we can generate more endocrine cells that we can study under the microscope and measure their physiological responses,” Chang-Graham said.
We have developed a system that enables us to study how the gut communicates with the rest of the body via whispered messages,” Hyser said.
“Our system has ‘raised the volume’ of the chemical whispers facilitating our investigations of the effects of the gut on health and disease and the testing of interventions to improve human health. For example, we can use our system to study how viral pathogens such as rotavirus regulate the responses of these endocrine gut populations and test strategies to modulate those responses.”

“The development of this model system also will allow researchers to begin investigating the complex interactions of the microbiome and diet with enteroendocrine cell function. Since several of the hormones and effector molecules released by enteroendocrine cells are already targets for the treatment of diseases such as type 2 diabetes and irritable bowel syndrome, we expect this model to aid in identifying novel therapeutics to treat these and other human diseases,” said co-corresponding author Dr. Robert Britton, professor of molecular virology and microbiology and member of the Alkek Center for Metagenomics and Microbiome Research and the Dan L Duncan Comprehensive Cancer Center at Baylor.
“This project was a revealing learning experience,” Chang-Graham said. “As the sole graduate student in the project, I had the opportunity to interact with postdocs and principal investigators directly and learn how collaborative science works and how to put together a paper like this.”
Find all the details of this work in the journal Cellular and Molecular Gastroenterology and Hepatology.
Other contributors of this work include Heather A. Danhof, Catherine Tomaro-Duchesneau and Umesh C. Karandikar (all from Baylor), and Melinda A. Engevik and James Versalovic (from Baylor and Texas Children’s Hospital).
Dr. Mary Estes is the Cullen Foundation Endowed Professor Chair of Human and Molecular Virology in molecular virology and microbiology at Baylor and emeritus founding director of the Texas Medical Center Digestive Diseases Center. She also is a member of the Dan L Duncan Comprehensive Cancer Center and holds a joint appointment in medicine–gastroenterology and hepatology at Baylor.
This project was supported by National Institutes of Health (grants F30 DK112563, U01 CA170930, R01 DK103759, U19 AI116497, R01 AI080656, R03 DK110270 and R01 DK115507), Baylor College of Medicine Seed Funding and Fonds de Recherche Santé Québec. This project also was supported in part by the National Institutes of Health PHS grant P30 DK056338 for the Texas Medical Center Digestive Diseases Center. Additional funding support for the Baylor College of Medicine Integrated Microscopy Core was provided by the National Institutes of Health (DK56338, CA125123), CPRIT (RP150578, RP170719), the Dan L Duncan Comprehensive Cancer Center and the John S. Dunn Gulf Coast Consortium for Chemical Genomics.