Chromosome remodeler SETD2 also remodels the cytoskeleton in kidney cancer cells
Kidney cancer begins when certain genes mutate and, as a result, alter how key molecules work. One of these key molecules is SETD2, which is known as a chromosome remodeler, that is, one that helps turn genes on in the nucleus of the cell. When researchers discovered that kidney cells that lose SETD2 become cancerous, all eyes turned toward SETD2 function in the nucleus of the cell to explain these cancers. It turns out that SETD2 not only remodels genes in the nucleus, it also remodels the cytoskeleton outside the nucleus.
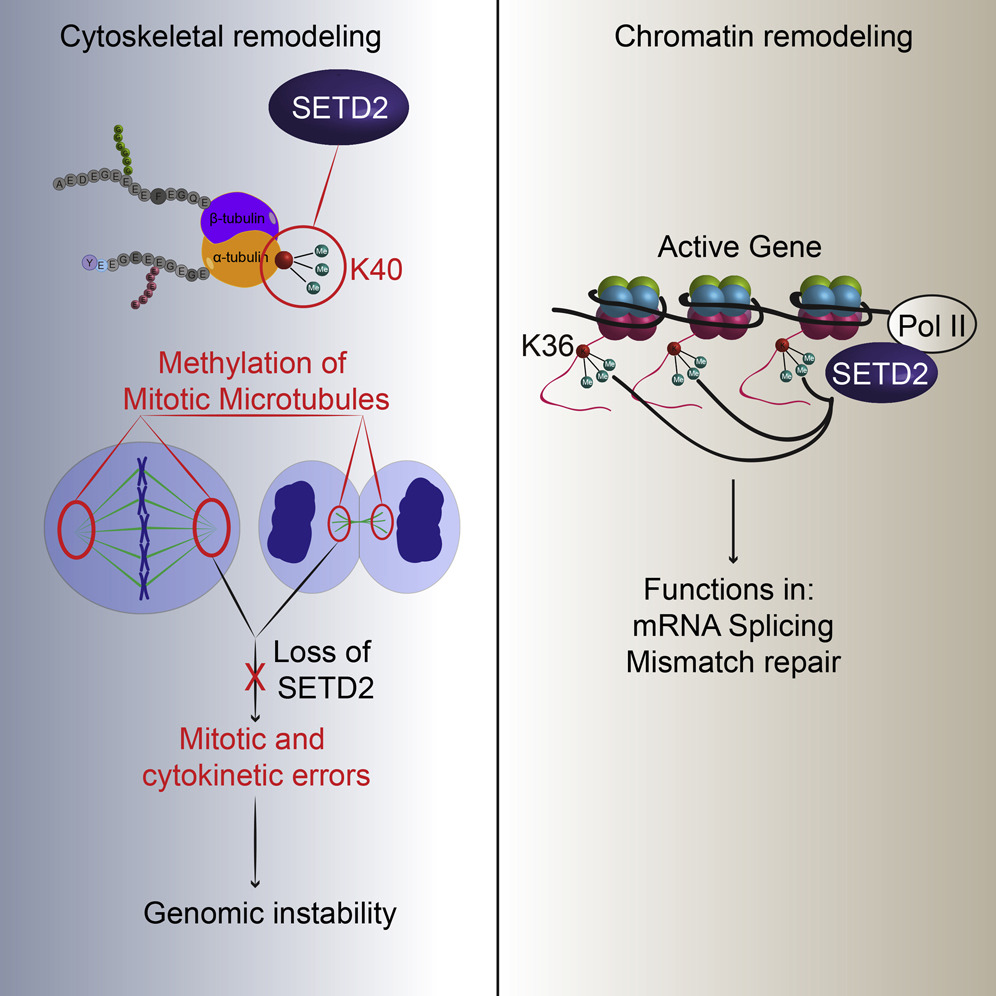
The cell’s cytoskeleton – a web of protein filaments and tubules – helps shape the cell, and much like our skeleton, also is used by cells for movement. Without SETD2, the cytoskeleton cannot help cells to properly move their chromosomes and divide. This discovery now opens new avenues for therapy focused on defects in the cytoskeleton, rather than on the nucleus.
“It was well known that SETD2 works as a chromosome remodeler by adding small chemical modifications that change chromosome structure to control gene expression. When SETD2 tags genes with a methyl group, this signals the activation of specific genes,” said senior author Dr. Cheryl L. Walker, director of the Center for Precision Environmental Health and professor of molecular and cellular biology at Baylor College of Medicine. Walker was the director of the Institute of Biosciences and Technology at Texas A&M University at the time the research was published.

“Our discovery of the new role of SETD2 on the cytoskeleton was serendipitous,” said first author Dr. In Young Park. “We were interested in understanding why kidney cells become cancer cells when they lose SETD2, so, like others in the field, we were comparing what happens to gene expression in cells that had SETD2 with what happens in cells that didn’t. We couldn’t find any cells available that did not have SETD2, so we genetically engineered a mouse model in which we could conditionally inactivate SETD2. We isolated embryonic fibroblasts from these mice and grew them in the lab. Unexpectedly, as soon as we triggered the fibroblasts to lose their SETD2, they behaved in a very peculiar way.”
The genetically engineered fibroblasts started the normal process of cell division by duplicating their DNA, but when it was time for each cell to separate its chromosomes and pull itself apart into two daughter cells, surprisingly, the cells couldn’t do it.
Under the microscope, as you can see in the video below, cells that had lost SETD2 (right side panel) struggled to separate into two cells, like two men trapped in a bag battling to get free,” said Walker. “Finally, the cells gave up trying to separate and remained as one cell with two nuclei. We saw this over and over again – these cells cannot figure out how to divide. This is not what we expected to see because cells cannot put a methyl mark on their genes.”
https://www.youtube.com/watch?v=E43L3sFGW4I
Embryonic fibroblast cells dividing into two daughter cells. Left, fibroblasts from normal mice complete cell division successfully dividing into two daughter cells. Right, fibroblasts from mice in which chromosome remodeler SETD2 has been inactivated. SETD2-deficient fibroblasts cannot divide into two daughter cells. (Courtesy of Walker lab)
This unanticipated observation started the researchers on a journey that led them to discover that SETD2 also tags the cytoskeleton with a methyl group to ensure the correct delivery of chromosomes and separation of daughter cells during cell division.
Researchers already knew that the cytoskeleton is tagged with chemical groups.
What is exciting about this work is that methylation has never been shown before to tag the cytoskeleton – the importance of this methyl tag to chromosome sorting and cell division was completely unexpected,” said Walker. “Using the same tag on both chromosomes, to control gene expression, and the cytoskeleton, to control movement, has never been seen before.”
“We know that there are many cancers that are defined by defects in chromosome remodelers such as SETD2, but everyone is looking at gene expression as the source of the problem,” said Walker. “Our results strongly suggest that there is at least a second source of a problem, the cytoskeleton, which controls movement, metastasis, and migration – very important functions for cancer cells.”
Without SETD2, the scientists think, critical cytoskeleton functions required for stability of the genome, to help the chromosomes move properly and dividing cells to separate into two daughter cells, are lost, promoting cancer development.
“Our results open a new road that can lead us to better understand and potentially successfully treat kidney cancer and the many other cancer types in which chromatin remodeler defects occur,” said Walker. “Importantly, this is not just a ‘one-off’ function for a chromatin remodeler. We have already discovered several others, in addition to SETD2, that play important roles in the cytoskeleton. It’s going to be a very exciting time for the field.”
Read all about this work in the journal Cell.
###
Other researchers who contributed to this work include Reid T. Powell, Durga Nand Tripathi, Ruhee Dere, Thai H. Ho, T. Lynne Blasius, Yun-Chen Chiang, Ian J. Davis, Catherine C. Fahey, Kathryn E. Hacker, Kristen J. Verhey, Mark T. Bedford, Eric Jonasch and W. Kimryn Rathmell. These researchers are affiliated with one or more of the following institutions: Texas A&M Health Science Center, Mayo Clinic Arizona, University of Michigan Medical School, University of North Carolina, The University of Texas MD Anderson Cancer Center and Vanderbilt University.
This work was supported in part by the Robert E. Welch Foundation (BE-0023), the National Institutes of Health (RC2ES018789, R01ES008263, R01ES023206 and P30ES023512) and the Cancer Prevention and Research Institute of Texas (CPRIT) (DP150086) grants R01GM070862, R01CA198482-01 and a V Foundation for Cancer Research award (T2012-008); R01CA166447; T32 GM008719 and NRSA award F30 CA192643-02; CPRIT RP110471; K12CA90628 and Gerstner Family Career Development Award and NIH/NCI grant P30CA016672.