PBRM1’s new function supports a novel cancer-promoting mechanism
When a protein is found mutated in a particular cancer, researchers look into the cellular location where the protein is typically present to investigate how its altered function can promote the disease. When mutations occur in chromatin remodelers, proteins that help turn genes on or off, scientists turn their eyes toward chromatin in the nucleus of the cell to explain these cancers.
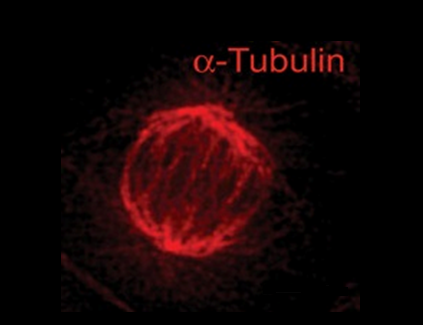
In 2016, Dr. Durga Nand Tripathi was part of the team in the lab of Dr. Cheryl Walker, director of the Center for Precision Environmental Health at Baylor College of Medicine, that made the unexpected discovery that chromatin remodeler SETD2 not only remodels chromatin in the nucleus, but also microtubules in the cytoskeleton outside the nucleus.

The cytoskeleton is a dynamic network of interlinking protein, thread-like structures, including filaments and microtubules that function as a “skeleton” for the cell. It gives a cell its shape and internal organization and provides mechanical support to carry out essential functions like cell division and movement.
The team discovered that, as it does for chromatin, SETD2 tags cytoskeleton microtubules with a methyl group. Loss of SETD2 resulted in defective segregation of chromosomes and problems with the separation of daughter cells during cell division. The findings suggested that defects in SETD2 could not only affect chromatin and change patterns of gene expression but also functions controlled by the cytoskeleton, such as proper cell division, movement and migration, which are important for cancer cells.
Methylation was not known to occur on microtubules,” said Tripathi, assistant professor in the Center for Precision Environmental Health, “so the cytoskeleton was not an obvious target for chromatin remodelers.”

SETD2 was only known to add methyl groups to histones, proteins that associate with DNA. The finding that SETD2 also can ‘write’ methyl groups on microtubules adds the new idea that cancer-associated defects in SETD2, and possibly other chromatin remodelers, could drive cancer development via their cytoskeletal activities.
Reading and writing methyl groups
Tagging histones with methyl groups, a known epigenetic mechanism, enables cells to regulate gene expression by turning specific genes on or off. The process involves both writing the methyl mark and “reading” it, which is accomplished by effector molecules such as transcription factors and large macromolecular complexes that can change the shape of chromatin or reposition nucleosomes to change chromatin accessibility.
“The epigenetic machinery responsible for establishing, modifying, and maintaining the system of epigenetic tags consists of proteins that work together to recognize (“read”), add (“write”) or remove (“erase”) marks such as methylation,” Tripathi said. “We proposed that since SETD2 was writing methyl marks on chromatin, the principle of reading these marks would also apply to the cytoskeleton. The mystery we set out to solve was, if SETD2 was writing methyl marks on microtubules, who was reading them?”

Tripathi and his colleagues, including his mentor Walker, as well as two postdoctoral fellows, Dr. Menuka Karki, specializing in microscopy, and Dr. Rahul Jangid, specializing in biochemistry, investigated whether the protein PBRM1 could be reading the methyl marks on microtubules.
Like SETD2, PBRM1 was frequently altered in many cancers, especially kidney cancers which also had defects in SETD2. However, based on their known chromatin functions, a connection between these two proteins could not be explained in terms of regulating gene expression.
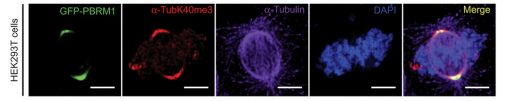
“We discovered that PBRM1 reads methyl marks on microtubules via a section of the molecule named BAH domain,” Tripathi said. “Importantly, previously another section of PBRM1 called the BD domain had been shown to read a different mark, an acetyl mark on histones of chromatin. But we found that, in addition to remodeling chromatin via the BD domain, PBRM1 also remodels the cytoskeleton via these BAH domains.”
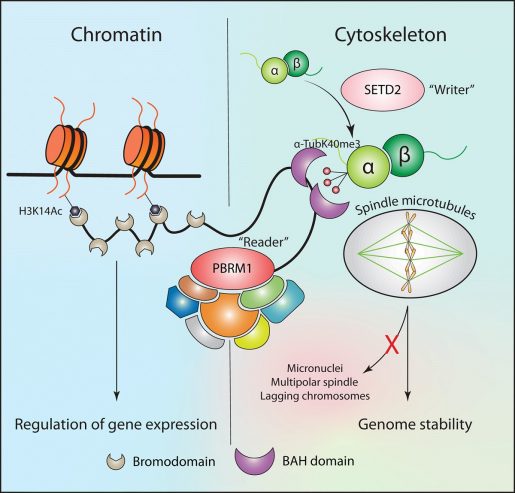
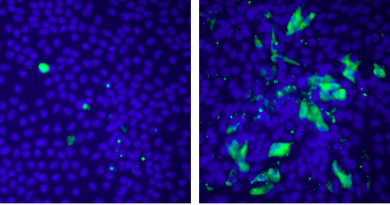
When the BAH domain is mutated, as occurs in kidney cancers, then the interaction between PBRM1 and the cytoskeleton is lost. This leads to the same type of cytoskeletal defects seen in cells that lose the SETD2 methyl mark: defects in chromosome segregation and genomic instability.
These findings suggest the new idea that cancer-associated defects in chromatin remodeler genes can drive cancer by disrupting cytoskeletal activities.
Interested in all the details of this work? Find them in the journal Science Advances.
Read the complete list of contributors as well as the financial support for this project in the publication.