The N-terminal domain of the androgen receptor is at the heart of its action
The androgen receptor binds to and mediates the effects of the androgenic hormone testosterone in both males and females. In both genders the androgen receptor regulates hair growth and sex drive. In males, it contributes to the development of sexual characteristics and, importantly, it drives the initiation and growth of prostate cancer, the second-leading cause of death in men.
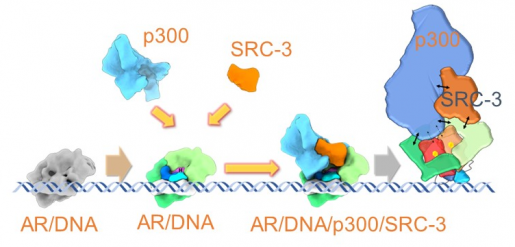

At Baylor College of Medicine, Dr. Ping Yi, assistant professor of molecular and cellular biology, and her colleagues strive to achieve a better understanding of how the androgen receptor works in cancer by looking in great detail at the 3-D structure of the receptor.
“Previous studies had shown that when the androgen receptor binds to testosterone, the resulting molecular complex travels to the cell nucleus where it interacts with DNA, turning genes on or off as necessary to regulate development and growth,” Yi said.
To carry on its activity, the androgen receptor also binds to coactivators and other molecules that promote its gene-regulatory function. However, how all these molecules are put together in a functional complex with the androgen receptor was not known.

“To treat prostate cancer effectively, we need to better understand how the androgen receptor works,” said co-corresponding author Dr. Bert O’Malley, chancellor and former long-time chair of the Department of Molecular and Cellular biology at Baylor. “In this study, we reveal for the first time the complete 3-D structure of the active, full-length androgen receptor-coactivator complex as it interacts with DNA.”
Cryo-EM reveals new components of androgen receptor structure and function
Before this study, researchers only had a partial idea of the 3-D structure of the androgen receptor. They were missing a part called the N-terminal domain, which biochemical evidence suggested might be crucial for its activity. Resolving the complete structure of the receptor would help understand why the N-terminal domain is key to the receptor’s activity.
“Cryo-electron microscopy let us see what the N-terminal domain of the androgen receptor looks like, how the protein is organized and how this and other individual domains contribute to the coactivated protein and its function,” said Yi, a member of Baylor’s Dan L Duncan Comprehensive Cancer Center.
The researchers discovered that the N-terminal domain at the beginning of the androgen receptor is where the coactivators bind, activating the complex that drives prostate cancer. This finding was in marked contrast with what the same researchers had discovered for the estrogen receptor, which is a major driver of breast cancer.
The androgen and the estrogen receptors belong to the same family of steroid nuclear receptors and share similar 3-D structures. However, despite having general structural similarities, in the estrogen receptor the coactivators bind not to the N-terminal domain at the beginning of the molecule, but to the C-terminal domain at the end of the molecule. This finding has important implications generating drugs for cancer treatment.
Therapeutic implications
“The androgen receptor drug inhibitors that are currently available for prostate cancer treatment bind to the C-terminal domain, which we found is not the main interactive site of the androgen receptor,” O’Malley said. “Our work strongly supports further studies to determine the effect that drugs directed at the androgen receptor’s N-terminal domain have on prostate cancer growth.”

“Our study provides a starting point to understand what is happening to the androgen receptor molecular machine in prostate cancer,” said co-corresponding author Dr. Zhao Wang, assistant professor in the Verna and Marrs McLean Department of Biochemistry and Molecular Biology. “Our findings also generate a broader therapeutic space for the treatment of not only prostate cancer but also related diseases, as well as new information on basic mechanisms of regulation of gene expression.”
Interested in learning all the details of this study? Find it in the journal Molecular Cell.
Other contributors to this work include co-first author Xinzhe Yu, Ross A. Hamilton, Hong Shen, Muyuan Chen, Charles E. Foulds, Michael A. Mancini and Steven J. Ludtke, all from Baylor College of Medicine.
This work is supported by NIH-NICHD and grants (HD8818, HD07857 and NIDDK59820, GM080139 and GM121203); DOD W81XWH-15-1-0536; CPRIT grant (RP150648); the Robert Welch Foundation (Q-1967-20180324 and BCM BMB department seed funds). Further support was provided by NCI Cancer Center Support Grant P30CA125123 (BCM Monoclonal Antibody/Integrated Microscopy Core/Recombinant Protein Expression Core Facility) and the Cryo-EM supported by Advanced Technology CPRIT Cryo-EM/ET Core (1RP190602) at BCM.