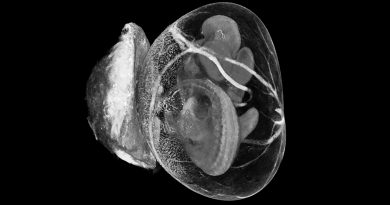
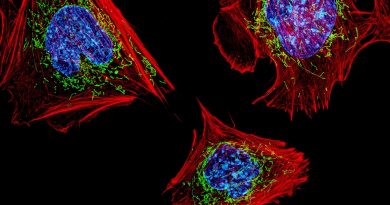
This map facilitates the search for epigenetic causes of disease
More than 15 years after scientists first mapped the human genome, most diseases still cannot be predicted based on one’s genes. Thus, researchers also are exploring epigenetic causes of disease, but these have been slow to come by. At Baylor College of Medicine, the lab of Dr. Robert A. Waterland has proposed a novel approach that can facilitate the study of epigenetics and disease.
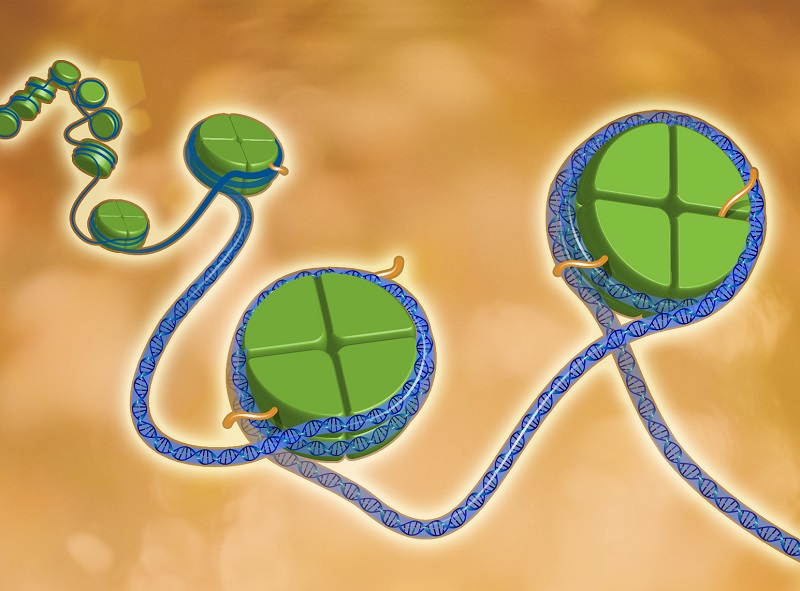
“Epigenetics is a system for molecular marking of DNA – it tells the different cells in the body which genes to turn on or off in that cell type. But the cell-specific nature of epigenetics makes it challenging to study,” said Waterland, professor of pediatrics – nutrition and of molecular and human genetics at Baylor. “Whereas a blood sample can be used to ‘genotype’ an individual, most epigenetic marks in blood DNA provide no clues about epigenetic dysregulation in other parts of the body, such as the brain or heart.”

The challenge
Waterland and his colleagues looked for regions of the genome that could be found in a blood sample and used to infer epigenetic regulation throughout the body, allowing scientists to test for epigenetic causes of disease.
To do this, they focused on the most stable form of epigenetic regulation – DNA methylation. The addition of methyl groups to the DNA molecule occurs in the embryonic state and can impact health for your entire life.
To identify genomic regions in which DNA methylation differs between people but is consistent across different tissues, they profiled DNA methylation throughout the genome in three tissues (thyroid, heart and brain) from each of 10 cadavers.
“Since these tissues each represent a different layer of the early embryo, we’re essentially going back in time to events that occurred during early embryonic development,” said Waterland, who also is professor at the USDA/ARS Children’s Nutrition Research Center and member of the Dan L Duncan Comprehensive Cancer Center. “To map DNA methylation, we converted methylation information into a genetic signal, then sequenced the genomes. Our atlas required massive amounts of sequencing data – 370 times more than were used for the first map of the human genome in 2001.”
The nearly 10,000 regions the researchers mapped out, called correlated regions of systemic interindividual variation (CoRSIVs), comprise a previously unrecognized level of molecular individuality in humans.
A novel way to uncover epigenetic causes of disease

“Recent studies are already showing that methylation at these regions is associated with a range of human diseases including obesity, cancer, autism, Alzheimer’s disease and cleft palate,” said Dr. Cristian Coarfa, associate professor of molecular and cellular biology, the Dan L Duncan Comprehensive Cancer Center and the Center for Precision Environmental Health at Baylor, and co-leader of the project
Waterland and his colleagues believe these findings will transform the study of epigenetics and disease, as researchers will now know where in the genome to look.
“Because epigenetic marking has the power to stably silence or stably activate genes, any disease that has a genetic basis could equally likely have an epigenetic basis,” Waterland said.
There is incredible potential for us to understand disease processes from an epigenetic perspective. CoRSIVs are the entryway to that.”
To read all the details of this study, go to the journal Genome Biology.
Other contributors to this work include Chathura J. Gunasekara, C. Anthony Scott, Eleonora Laritsky, Maria S. Baker, Harry MacKay, Jack D. Duryea, Noah J. Kessler, Garrett Hellenthal, Alexis C. Wood, Kelly R. Hodges, Manisha Gandhi, Amy B. Hair, Matt J. Silver, Sophie E. Moore, Andrew M. Prentice, Yumei Li, and Rui Chen.
See a list of author affiliations.
This work was supported by the Bill and Melinda Gates Foundation, NIH/NIDDK (1R01DK111522), the Cancer Prevention and Research Institute of Texas (RP170295), USDA/ARS (CRIS 3092-5-001-059), UK Medical Research Council (MC-A760-5QX00), The Bill and Melinda Gates Foundation (OPP1 066947), NIH (S10OD023469), NIH/NICHD (1R21HD087860) and Wellcome Trust (098386/Z/12/Z).