Not all human norovirus strains are created equal, in their sensitivity to interferon
Human noroviruses (HuNoVs) cause the majority of the cases of viral gastroenteritis in the world and bring about significant mortality in all age groups; yet, there are still no vaccines or other approved therapeutic strategies available. One reason for the limited options to prevent and treat this disease is the difficulty in growing the virus in the lab to understand how the virus makes people sick.
“In-depth studies of how virus and host interact have been possible only recently thanks to the development of several laboratory cultivation systems,” said first author Shih-Ching Lin, a graduate student in the laboratory of Dr. Mary Estes at Baylor College of Medicine. “In this study, we worked with human intestinal enteroids (HIEs), a cultivation system that recapitulates many of the characteristics of the human infection, to investigate their response to HuNoV infection and how it affects viral replication.”
New findings illuminate the interplay between HuNoV and HIEs
The researchers infected HIEs with HuNoV strain GII.3 or pandemic strain GII.4 and determined which HIE genes were activated as a result.

“We discovered that both strains preferentially triggered a type III IFN response, including activation of a number of IFN-stimulated genes, as well as increases in a subset of long non-coding RNAs. Changes in long non-coding RNAs, which are known to regulate gene expression, had never been reported before for gastrointestinal virus infections,” Lin said.
Next, Lin, Estes and their colleagues studied the effect of IFN on the replication of HuNoV. Adding IFN to the cultures reduced replication of both strains, suggesting that IFN may have value as therapeutics for HuNoV infections. This could be important for chronically infected immunocompromised patients who can suffer with diarrhea for years.
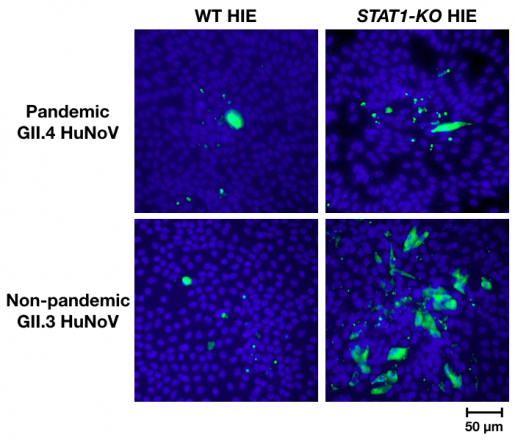
To obtain insights into the genes of the IFN pathway that contribute to the antiviral response to HuNoV, the researchers knocked out several of these genes in HIE cultures, infected them with strain GII.3 or GII.4 and measured the rate of viral proliferation.
We expected that the absence of IFN responses by HIEs would promote viral replication in both strains. It was surprising and very exciting to find significant strain differences,” Lin said.
“We saw that only strain GII.3 was able to spread and multiply more when HIEs could not activate IFN responses. Replication of GII.4, on the other hand, was not enhanced,” said Estes, Cullen Foundation Endowed Chair and Distinguished Service Professor of molecular virology and microbiology at Baylor. Estes also is a member of the Dan L Duncan Comprehensive Cancer Center. “It was exciting to see the GII.3 strain proliferate and spread in the cultures as we had never seen it before.”

“The strain-specific sensitivities of innate IFN responses to HuNoV replication we observed provide a potential explanation for why GII.4 infections are more widespread and become pandemic,” Lin said. “Our findings also show the importance of keeping potential strain differences in mind when studying the biology of HuNoV and developing therapies. Our new genetically modified HIE cultures also will be useful tools for studying innate immune responses to other viral or microbial pathogens.”
Find all the details about this work in the Proceedings of the National Academy of Sciences USA.
Other contributors to this work include Lin Qu, Khalil Ettayebi, Sue Crawford, Sarah Blutt, Matthew Robertson, Xi-Lei Zeng, Victoria Tenge, B. Vijayalakshmi Ayyar, Umesh Karandikar, Xiaomin Yu, Cristian Coarfa, Robert Atmar and Sasirekha Ramani, all at Baylor College of Medicine.
This work was supported in part by Public Health Service grants (PO1AI057788, U19AI144297, P30DK56338), T32DK07664 and contract HHSN2722017000381 from the National Institutes of Health. Additional support was provided by the Cancer Prevention Institute of Texas (RP160283), Baylor College of Medicine Comprehensive Cancer Center Training Program and RP17005, NIH P30 shared resource grant CA125123, NIEHS grants P30ES030285 and P42ES0327725 and the John S. Dunn Research Foundation.