Regulating gene regulatory regions offers a novel mechanism to control ATXN1 levels
People with a rare genetic neurodegenerative disease known as spinocerebellar ataxia type 1 (SCA1) usually have progressive problems with movement, including loss of coordination, and balance (ataxia) and muscle weakness. They typically survive 15 to 20 years after symptoms first appear.
“SCA1 is one of the adult-onset neurodegenerative diseases for which we know the genetic cause, in this case the gene ATXN1,” said first author Larissa Nitschke, doctoral candidate in the lab of Dr. Huda Zoghbi at Baylor College of Medicine and the Jan and Dan Duncan Neurological Research Institute at Texas Children’s Hospital. “When we identified the gene, we learned that mutations can cause the ATXN1 protein to remain in cells longer than normally. This is bad news for neurons as too much ATXN1 leads to their death.”

The findings suggested that lowering the levels of ATXN1 might result in improved symptoms, so Nitschke and her colleagues looked for mechanisms that cells use to control the levels of ATXN1.
How cells regulate ATXN1 levels
As with other genes, part of the ATXN1 gene codes for the protein itself and the rest is involved in regulating the expression of the RNA and protein encoded by the gene.
“We looked at a regulatory region known as 5-prime untranslated region (5’ UTR), which is unusually long for the ATXN1 gene, and found that it keeps the protein in check so it does not accumulate to reach toxic levels,” Nitschke said.
The researchers studied this region in great detail, piece by piece, looking to identify individual sequences or elements that might control the amount of ATXN1 that cells produce. They found several elements that fulfilled that function.
Nitschke and her colleagues focused on one regulatory element that seemed important because it is conserved in many species. They discovered that this short piece could regulate ATXN1 levels.
We also found that we could reduce the amount of ATXN1 produced with a microRNA called miR760 that binds specifically to the conserved small piece in the 5’UTR region.”
“MicroRNAs are tiny RNA molecules that cells use to regulate the production of specific proteins by interacting with regulatory regions,” Nitschke said. “This finding encouraged us to test whether miR760 could reduce the amount of ATXN1 in animal models of SCA1.”
Reducing ATXN1 in the cerebellum improves SCA1 symptoms in animal models
Testing the effect of miR760 on animal models of SCA1 had to be planned carefully.

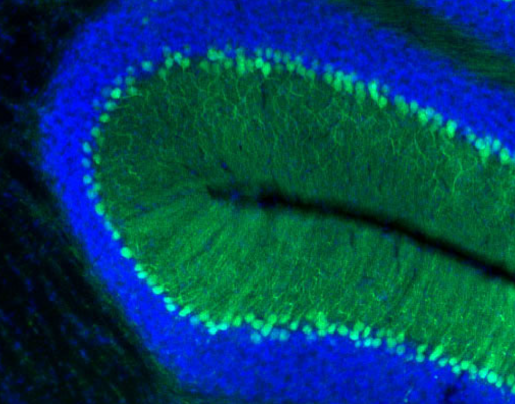
“The role of ATXN1 in the brain is complex,” said Zoghbi, professor of molecular and human genetics, pediatrics and neuroscience, and Ralph D. Feigin, M.D. Endowed Chair at Baylor. “Having too much ATXN1 in the back of the brain, the region called the cerebellum, which is involved in balance and coordination, results in balance problems. Having too little ATXN1 in the part of the brain for learning and memory increases the risk of Alzheimer’s disease.”
The researchers designed their experiments to reduce the levels of ATXN1 only in the cerebellum using gene therapy directed just at this brain region. The results were encouraging. Providing miR760 lowered the levels of ATXN1 and, importantly, improved motor and coordination deficits in the animal models of SCA1.
“The most exciting part of our findings was that we could reduce some of the symptoms of SCA1 in the animal models,” Nitschke said. “Although we only lowered the levels of ATXN1 by about 25 percent, the mice significantly improved their movements. This result strongly supports further studies to explore the effectiveness of this approach to treat the human condition.”
The findings not only highlight the importance of ATXN1 gene regulatory regions in SCA1, but also bring up the possibility that mutations in these DNA elements could lead to increased levels of ATXN1 and in turn increase the risk for balance problems,” said Zoghbi, director of the Jan and Dan Duncan Neurological Research Institute and member of the Howard Hughes Medical Institute and the Dan L Duncan Comprehensive Cancer Center. “Identifying and analyzing the sequences of such elements in people with balance problems might have a potential to help provide a diagnosis.”
Are you interested in reading the complete study? Find it in the journal Genes & Development.
Other contributors to this work include Ambika Tewari, Stephanie L. Coffin, Eder Xhako, Kaifang Pang, Vincenzo A. Gennarino, Jennifer L. Johnson, Francisco A. Blanco and Zhandong Liu. The authors are associated with one or more of the following institutions: Baylor College of Medicine, Jan and Dan Duncan Neurological Research Institute at Texas Children’s Hospital and Howard Hughes Medical Institute, Houston.
This work was funded by NINDS grant 4R37NS027699 (HYZ), the Howard Hughes Medical Institute, Baylor Research Advocates for Student Scientists, the Young Investigator Research Grant by the National Ataxia Foundation and the IDDRC at Baylor College of Medicine grant 1U54HD083092 (Administrative and Behavioral Cores) from the Eunice Kennedy Shriver National Institute of Child Health & Human Development. Further support was provided by the Gene Vector Core at Baylor College of Medicine.