Research opens the door to improved treatments for congenital heart disease
Researchers at Baylor College of Medicine, Texas Heart Institute, Texas Children’s Hospital and collaborating institutions have improved our understanding of the mechanisms underlying the progression of congenital heart disease (CHD) – a spectrum of heart defects that develop before birth and remain one of the leading causes of childhood death.
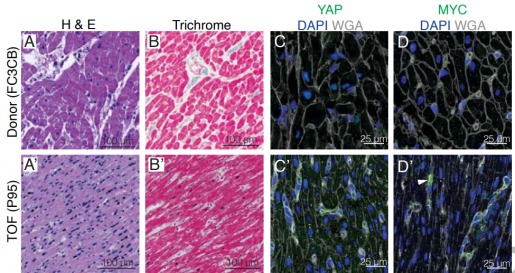
The research published in Nature represents the first reported single cell analysis of the genes expressed by heart cells and immune system cells of congenital heart disease patients. The team uncovered key differences among different types of CHD, which provides insights into the mechanisms of the disease, helps define clinical outcomes and illuminates opportunities to devise personalized treatments.

“When the parents of a child with a newly diagnosed congenital heart disease approach me, they always ask me two questions: ‘Why does my child have this problem?’ and ‘What are you going to do about it?’” said co-author Dr. Diwakar Turaga, assistant professor of pediatrics at Baylor and a pediatric cardiac critical care specialist at Texas Children’s. “This study addresses both questions.”
To identify the genetic underpinnings of CHD, the researchers obtained heart tissue samples from multiple patients, covering different forms of the condition.
They analyzed the gene expression signature of single cells using a technique called single nuclear RNA sequencing (snRNA-seq). The team studied different types of heart cells and immune cells in each tissue sample. The analysis revealed genetic pathways that are activated in each cell, a way of knowing what the cells are doing.
“This unprecedented analysis showed what abnormal genetic changes each patient has, answering the first question about what causes the condition,” Turaga said. “Having this information opens the door to answer the second question. The alterations at the genetic and cellular levels that lead to the abnormal development of the heart guide the researchers and physicians’ decisions on how to treat CHD.”

Our comprehensive analysis showed us something new – to expect genetic alterations not only in the heart, but in immune cells as well.
“Therefore, treating CHD requires a multiprong approach. It involves both repairing the genetic pathways that are altered in heart cells and modulating the damaging pro-inflammatory activity of immune system cells,” said corresponding author Dr. James Martin, vice chair and professor of integrative physiology, professor and Vivian L. Smith Chair in Regenerative Medicine at Baylor and director of the Cardiomyocyte Renewal Lab at the Texas Heart Institute.
Our goal is a better understanding of the biology of CHD, but as we move forward, we also would like to develop personalized treatments for these patients.
There is still a lot of work to do as the team, including co-first author Dr. Matthew C. Hill, heads toward that goal. This is really the beginning to a personalized medicine approach to treat these conditions.”

“Patients with congenital heart disease are at a much greater risk of heart failure than the general population, despite the amazing surgeries performed following birth to correct their intrinsic heart defects,” said co-first author Zachary Kadow, a student in the Medical Scientist Training Program (M.D./Ph.D.) at Baylor. “We discovered genetic pathways activated in the congenital heart disease that are different from those in adult heart failure. This work is a start toward our long-term goal of developing therapies that specifically slow heart failure progression in congenital heart disease patients, allowing them to live longer, healthier lives.”

“The most exciting part of this work was the diverse range of congenital heart disease covered, in addition to the novel technologies used. Our findings provide important biological insights into the cellular and molecular dynamics in congenital heart disease and will have significant implications for both diagnostic and therapeutic development,” said co-author Hali Long, graduate student in the Martin lab.

“It’s very exciting that this is a product of a powerful collaboration between one of the most sophisticated laboratories in cardiovascular research and the top pediatric heart center,” said co-author Dr. Iki Adachi, associate professor of surgery at Baylor and Clayton Endowed Chair in Cardiac Transplant and Mechanical Support at Texas Children’s.
Other contributors to this work include Yuka Morikawa, Thomas J. Martin, Emma J. Birks, Kenneth S. Campbell, Jeanne Nerbonne, Kory Lavine, Lalita Wadhwa and Jun Wang. The authors are affiliated with Baylor College of Medicine, Texas Heart Institute, Texas Children’s Hospital, The University of Texas Health Science Center at Houston, University of Kentucky and Washington University School of Medicine, St. Louis.
This work was supported by the Department of Defense (CDMRP) (W81XWH-16- PRMRP-IIRA), National Institutes of Health (1F31HL156681-01, F30HL145908, 5T32HL007208-42, R56 HL142704, R01HL142704, R01HL 127717, R01HL 130804 and R01HL 118761) and the Vivian L. Smith Foundation. Further support was provided by Baylor Research Advocates for Student Scientists and Baylor College of Medicine Medical Scientist Training Program, LeDucq Foundation’s Transatlantic Networks of Excellence in Cardiovascular Research (14CVD01: “Defining the Genomic Topology of Atrial Fibrillation”), the MacDonald Research Fund Award (16RDM001), a grant from the Saving Tiny Hearts Society, NIH HL149164, HL148785 and University of Kentucky Myocardial Recovery Alliance. The TCBR is supported by Children’s Discovery Institute of Washington University and St. Louis Children’s Hospital (PM-LI-2019- 829), NIH through M. D. Anderson’s Cancer Center Support Grant CA016672, the NCI’s Research Specialist 1 R50 CA243707-01A1, and a Shared Instrumentation Award from the Cancer Prevention Research Institution of Texas (CPRIT) (RP121010 and RP180672) and NIH (CA125123 and RR024574, 1S10OD023469).