A faster, less expensive approach identifies highly specific anti-cancer compounds
Bromodomain (BD)-containing proteins are implicated in cancer, inflammation, infectious diseases and metabolic disorders and have emerged as potential drug targets in a variety of diseases.

“Over a decade of research has shown that BD-inhibitors can help control cancer growth; however, when tested in clinical trials some had side effects and limited efficacy, halting further clinical development. This has encouraged our group to search for more effective BD-inhibitors,” said Dr. Joanna Yi, assistant professor of pediatric hematology/oncology at Baylor College of Medicine and Texas Children’s Hospital, and one of the lead authors of this work.”

Searching for highly-specific anti-cancer compounds
The researchers focused on identifying inhibitors specific for the first bromodomains (BD1) in the bromodomain and extra-terminal (BET) subgroup of human proteins. Recent research has shown that BD1 is very important in driving cancer, the researchers explained.

“To identify novel BD1-inhibitors, we took advantage of an innovative, faster and more cost-effective drug discovery tool called DNA-Encoded Chemistry Technology (DEC-Tec) developed at Baylor’s Center for Drug Discovery (CDD), which enables us to screen billions of compounds,” said first author Dr. Ram K. Modukuri, a staff scientist in the Department of Pathology & Immunology and the CDD at Baylor.
The more commonly used method for discovering drugs, called high throughput screening, involves screening at most a million compounds in individual test tubes. In contrast, by using DEC-Tec, the team was able to screen 4 billion DNA-encoded molecules all in one test tube against BD1 to find one that would bind to it with high specificity when compared to binding to other bromodomains.
“DEC-Tec enabled us to identify CDD-724, a compound that is highly selective for BD1. It is about 2,000 times better at inhibiting BD1 than at inhibiting other human bromodomains, including the second bromodomain (BD2) of the BET subgroup,” Modukuri said.

How does DNA-Encoded Chemistry Technology work?
“The DNA-Encoded Chemistry Technology process involves a simultaneous screen of billions of molecules, each one tagged with a DNA barcode,” said corresponding author Dr. Martin Matzuk, professor and chair of pathology and immunology and director of the Center for Drug Discovery at Baylor. “The molecules that ‘stick’ to the protein (in this case BD1) are identified by sequencing of their attached DNA barcode.
This is a rapid drug discovery screen, and our study demonstrates its enormous potential to find unique candidates for anti-cancer drugs.”

To better understand why their BD1-inhibitor stands out over other inhibitors, the team joined forces with Dr. Choel Kim, associate professor of pharmacology and chemical biology, who also is a member of the CDD and the Dan L Duncan Comprehensive Cancer Center at Baylor. The researchers conducted 3D molecular studies to determine the precise location on the BD1 protein to which the BD1-inhibitor binds. They discovered that the BD1-inhibitor binds to a shallow area on the BD1 protein – this is not seen in other BD1 inhibitors. This finding represents a new opportunity to explore other BD1-selective inhibitors.
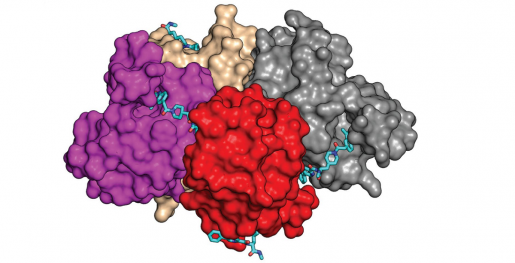
“We are searching for highly specific, potent, effective compounds with reduced side effects that we can bring to the clinic,” said Yi, also a member of Baylor’s CDD and the Dan L Duncan Comprehensive Cancer Center. “We are ready to test these compounds in animal models to evaluate their safety and efficacy, which is one step closer to clinical trials.”
Read all the details in the Proceedings of the National Academy of Sciences.
Zhifeng Yu, Zhi Tan, Hai Minh Ta, Melek Nihan Ucisik, Zhuang Jin, Justin L. Anglin, Kiran L. Sharma, Pranavanand Nyshadham, Feng Li, Kevin Riehle, John C. Faver, Kevin Duong, Sureshbabu Nagarajan, Nicholas Simmons, Stephen S. Palmer, Mingxing Teng and Damian W. Young also contributed to this work. The authors are affiliated with Baylor College of Medicine and/or Texas Children’s Hospital.
This work is supported by the Bill and Melinda Gates Foundation (INV-001902), the Eunice Kennedy Shriver National Institute of Child Health and Human Development (P01HD087157), the Welch Foundation (H-Q-0042) and a Core Facility Support Award from the Cancer Prevention Research Institute of Texas (CPRIT) (RP160805). Further support was provided by NIH grants (5K12CA090433-17, R01DK121970, R61HD099995, S10RR25528, S10RR028976, and S10OD027000), Alex’s Lemonade Stand Foundation, the Curing Kids Cancer Foundation, CURE Childhood Cancer Foundation, and CPRIT grants (RR220012 and RR220039).