Research Spotlight: Dr. Fritz Sedlazeck assists in filling gaps in human genome, sequencing Rice University owl genome and more
Dr. Fritz Sedlazeck, associate professor in the Human Genome Sequencing Center at Baylor College of Medicine, is passionate about his genomics research, which includes tools he has developed to identify structural variations (SV) – changes in the genome that can alter the regulation of gene expression and impact human disease. Those same SV tools were useful for quality control in the final push to complete the last 8% of the first human reference genome, originally sequenced in 2000.

Contributing to the completion of the first human genome
The missing 8% of first human reference genome spanned gaps in multiple areas, including the telomeres, those end caps of a chromosome that are filled with repetitive data and serve as a kind of buffer to protect the rest of the chromosome. The group of researchers striving to fill the genome gaps were known as the Telomere-to-Telomere (T2T) Consortium, and they announced completion of the sequencing on March 31, 2022.

“Some of my former colleagues from Johns Hopkins were among the T2T Consortium, and my identity was known to others due to my SV modeling tools,” Sedlazeck said. “Although my models may have supported only the quality control team, I certainly felt the excitement rippling across the entire group as the consortium neared completion of that 3.055 billion base pair sequence.”
He might be modest about his contributions, but Sedlazeck is affiliated with two studies published in the journal Science in March and April by the T2T Consortium about the now complete human genome sequence.
Helping to sequence the genome of Rice University’s mascot
Several months before the T2T genome sequence was published, Sedlazeck had been invited to collaborate with Rice University and the Houston Zoo to sequence a great horned owl genome.
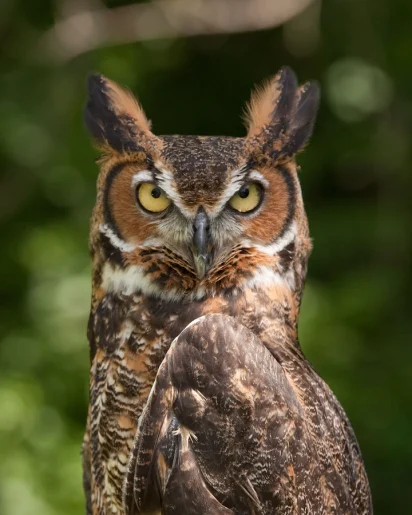

“Over recent years, I have been active in helping people to assemble a genome of their interest and quality control the outcomes,” Sedlazeck said. “The horned owl genome struck me as compelling because there are interesting microchromosomes to study. In bird genomes, larger chromosomes and many smaller microchromosomes co-exist; mammal genomes do not have microchromosomes.”
Dr. Todd Treangen — one of Sedlazeck’s collaborators at Rice — worked with officials at the Houston Zoo for three years to meet requirements for obtaining a sample and transferring it from the Zoo to Baylor’s Human Genome Sequencing Center lab. There, Sedlazeck extracted strings of DNA from the blood sample and performed quality control. He then set up the DNA to run through the lab’s PacBio Sequel II, which could produce long-read sequencing.
Long-read sequencing was key to the T2T Consortium’s work in mapping the final 8% of their human genome sequence, and it was equally important for the robust and rapid assembly Treangen had in mind for the great horned owl sequence.”
In mid-January, Sedlazeck shared a ZIP file of 10-15,000 base pairs with Treangen, who was recruiting a team of students for the next phase of the process. One of Rice collaborators, Dr. Genevera Allen, is the founder and faculty director of the Data to Knowledge (D2K) Lab. Two D2K undergraduate students and one Ph.D. student were matched with Treangen’s group for their capstone project: assembling the first great horned owl genome sequence.
‘Teaching is an essential part of being a scientist’
“Another factor of the owl genome project that appealed to me was indeed the prospect to take this material to students, to give them a better sense about computational biology and genomics and hopefully just have some fun with data exploration,” Sedlazeck said. “I hope that the data produced here at Baylor and the work done together with Rice will enable new insights into the genomic complexity of this owl. Nevertheless, I think it is more important that this year’s students obtain insights into genomics and computational biology and maybe we sparked interest for these fascinating fields.”
The most complete owl genome sequencing (prior to the Rice collaboration) was for a different species, published in 2017. Sedlazeck said his work with the Rice group demonstrated that the maturity of the sequencing technology (PacBio) and algorithms have come a long way in just five years.
Like the ripples of excitement he felt as part of the T2T sequencing efforts, Sedlazeck sensed the Rice students’ growing delight with their progress.
“Science is serious enough; it is good to have fun with it every now and then,” he said.
I think it is important to invest time and resources in projects like these, so that students experience more hands-on learning about the current state of algorithms and the algorithmic limitations, to spark their interest and also to have fresh minds thinking about an ‘old’ problem.”
“Thus, teaching students and leading workshops is something that I truly enjoy and is — I think — an essential part of being a scientist. What good are our results if we cannot communicate or fascinate people with them?”
Research that improves medical diagnosis
Although he thoroughly enjoyed the owl genome side project and working with Rice and the Houston Zoo, Sedlazeck is anxious to return to his work on human genomes. His sequencing work may help differentiate variables across human genomes and infer markers for heart attacks and other diseases.
One of the lab’s collaborations with the National Institute of Standards and Technology (NIST) helped develop benchmarks for medically relevant genes and how they could be assessed or characterized. Utilizing current technologies and techniques, Baylor and NIST developed software benchmarks that improved the scientists’ understanding of genes with critical implications for health.
But one of the breakthroughs Sedlazeck finds even more exciting is being part of the first group to achieve sequencing and diagnosis within 7-8 hours of drawing a sample. Both patients and their physicians will benefit from this rapid turn-around. Currently, a patient with an undiagnosed disease meets with their doctor, who requests a blood sample be taken and sent to a lab for sequencing. Standard blood sample sequencing usually takes weeks or even months. For the parents and physicians of newborns dealing with undiagnosed illnesses and suffering life-threatening symptoms, even the 14-20 hours for a priority sequencing feels too long.
“I worked with a team at Stanford University that developed software and lab methodology, called Ultrarapid Nanopore Genome Sequencing, to analyze blood and diagnose disease markers with high accuracy and speed,” Sedlazeck said. “Within 8 hours, or possibly 7 hours, from the time the sample is drawn, we can hand off a clinical diagnostic report to the physician.”
That is a Guinness World Record! The T2T work also was recognized as a Guinness World Record for the most complete human genome.”
Contributed by Carlyn Chatfield, Rice University