Wnt signaling is identified as a target in NEDAMSS disorder
A recent study conducted by researchers at Baylor College of Medicine and Texas Children’s Hospital is the first to identify an underlying mechanism and possible drug targets for a new severe neurological disorder in children called NEDAMSS.
The disorder, called NEDAMSS for neurodevelopmental disorder with regression, abnormal movements, loss of speech and seizures, is caused by spontaneous mutations in the Interferon Regulatory Factor 2 Binding Protein Like (IRF2BPL) gene. In this study, the team led by Drs. Hugo Bellen, Paul Marcogliese and Debdeep Dutta at Baylor and the Jan and Dan Duncan Neurological Research Institute (Duncan NRI) at Texas Children’s employed fruit flies to dissect the biological function of IRF2BPL.
In 2018, Marcogliese and Bellen, along with a team of researchers affiliated with the Undiagnosed Diseases Network – a National Institutes of Health-funded network whose goal is to find genetic variants responsible for previously unidentified disorders – discovered the NEDAMSS disorder. In collaboration with Drs. Shinya Yamamoto, Michael Wangler and others at the Duncan NRI and Baylor, as well as Drs. Loren Pena and Vandana Shashi at the UDN’s Duke clinical site, they identified mutations in IRF2BPL as the cause of severe steady regression of motor and language skills in the initial cohort of five patients.

“Today, there are 31 patients identified with mutations in IRF2BPL,” said Marcogliese, co-corresponding author of the study and postdoctoral associate in the Bellen lab. “IRF2BPL has been named as ‘one of the top 100’ autism candidate genes and its variants have been implicated in early-onset Parkinsonism. However, until now, very little was known about the biological function of this gene and how its loss results in neuronal dysfunction.”
IRF2BPL/Pits act via the Wnt/Wg pathway
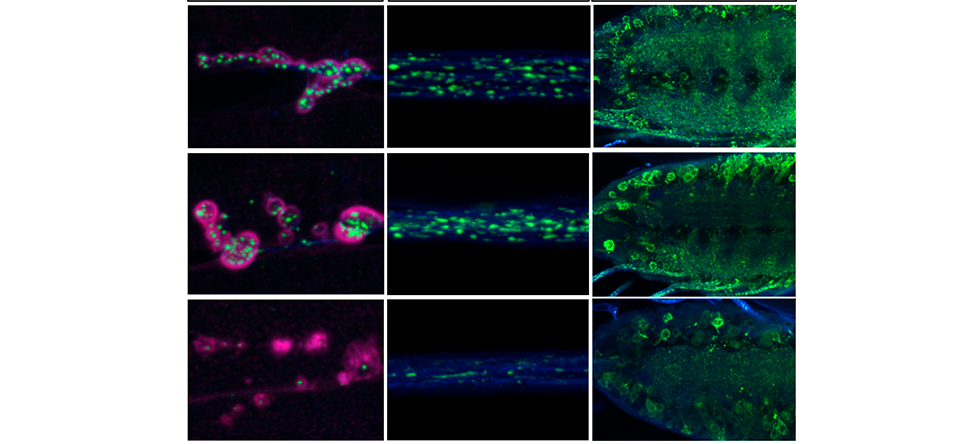
Interestingly, overexpression of Pits (the fly version of IRF2BPL) or human IRF2BPL in flies resulted in serrated wings and bristle loss, both of which are common when the Wnt/Wingless signaling pathway is perturbed during development. Reducing Pits in neurons, however, led to an age-dependent neurological defect in flies, which the team later found was associated with an increase in the activity of the Wnt/Wingless pathway.
In collaboration with Dr. Nan Cher Yeo at the University of Alabama – Birmingham and Dr. Kathrin Meyer at Nationwide Children’s Hospital, they confirmed these findings in zebrafish and in patient-derived cells as well.
Numerous experimental observations led the researchers to conclude that Pits and Wnt/Wg act in an antagonistic manner. In agreement with these observations in animal models, they found that Wnt signaling is increased in NEDAMSS patients, whose cells are known to have reduced levels of IRF2BPL.
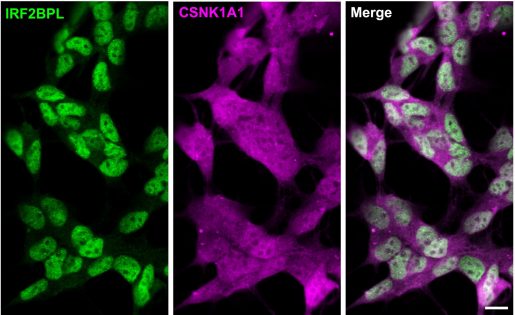
To understand how Pits/IRF2BPL regulates Wg/Wnt pathway, the researchers conducted biochemical and mass spectrometry assays, which revealed a Wnt antagonist, Casein kinaseIα (CkIα), as one of the top binding partners of Pits. Further, they also confirmed genetic interactions between IRF2BPL and CkIα.
In summary, the study shows an antagonistic relationship between IRF2BPL/Pits and the Wnt/Wg pathway in the fruit fly model. It also shows that under normal circumstances, IRF2BPL/Pits regulates neural function by inhibiting the Wnt/Wg pathway.
In NEDAMSS patients, loss of IRF2BPL is accompanied by increased levels of Wnt in astrocytes – a prominent type of cell in the brain. The Wnt pathway, the human counterpart of the Wingless pathway in flies, is known to be important for the proper development and function of neurons and other organs, and its disruption results in several types of cancers. The researchers also showed that they can suppress characteristics associated with the loss of IRF2BPL/Pits using Wnt inhibitors, which have proven to be safe and effective in treating many cancers.
It used to take decades from the initial discovery of a novel gene to dissecting its biological mechanism to finding a therapy,” said Bellen, Distinguished Service Professor of Molecular and Human Genetics at Baylor and the corresponding author of the work. “However, with the UDN’s unique approach that fosters close collaborations between clinicians and researchers working on various model systems, we were able to move from the initial identification of this mutation to a potential therapy in just three years.”
“Furthermore, we were able to progress rapidly thanks to the continued support from the iDREAM For a Cure, as well as supplemental support from the NIH for which we are truly grateful. We hope to translate these promising discoveries into a viable therapy in the future.”
The researchers report their findings in the journal Science Advances.
Other researchers involved in the study include Shrestha Sinha, Nghi Dang, Zhongyuan Zuo, Yuchun Wang, Di Lu, Fatima Fazal, Thomas Ravenscroft, Hyunglok Chung, Oguz Kanca, JiJuWan, Emilie Douine, Stanley Nelson, Matthew Might, Kathrin Meyer and Nan Cher Yeo. They are affiliated with one or more of the following institutions: Baylor College of Medicine, Texas Children’s Hospital, the Research Institute at Nationwide Children’s Hospital, University of Alabama, David Geffen School of Medicine, Cincinnati Children’s Hospital and Ohio State University.
Read more.