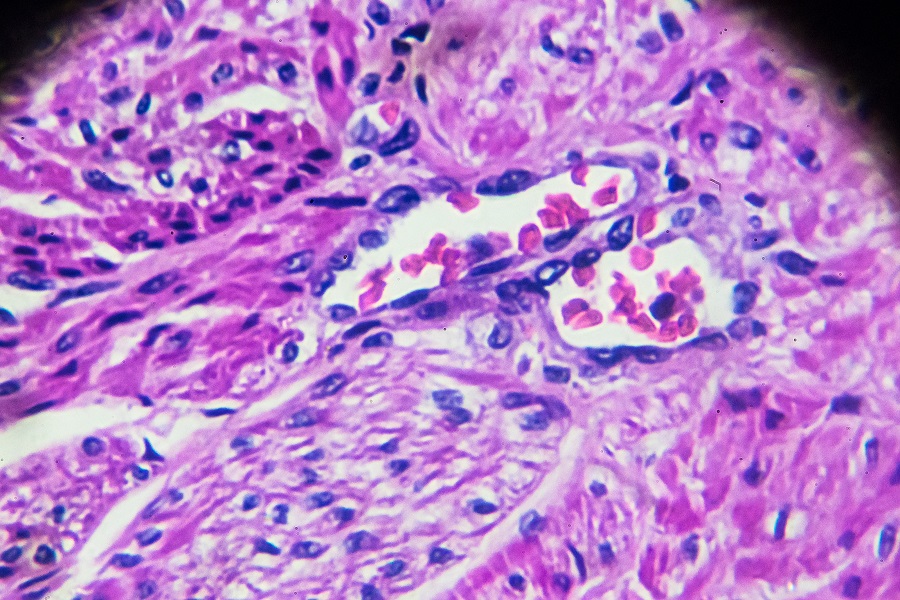
Cells arrange their chromosomes following one of two designs
A team of biologists studying the tree of life has unveiled a new classification system for cell nuclei according to how chromosomes, which can be several meters long, fold up to fit inside the nuclei. They also discovered a method for transmuting one type of cell nucleus into another.

The study, which appears in the journal Science, emerged from several once-separate efforts. One effort is centered on the DNA Zoo, an international consortium spanning dozens of institutions including Baylor College of Medicine, the National Science Foundation-supported Center for Theoretical Biological Physics (CTBP) at Rice University, the University of Western Australia and SeaWorld. The other effort emerged from an independent team in the Netherlands Cancer Institute.
Cells arrange their chromosomes following one of two patterns

“Whether we were looking at worms or urchins, sea squirts or coral, we kept seeing the same folding patterns coming up,” said Dr. Olga Dudchenko, co-first author of the new study and a member of the Center for Genome Architecture at Baylor and CTBP.
Eventually, the team realized it was just seeing variants on two overall nuclear designs. “In some species, chromosomes are organized like the pages of a printed newspaper, with the outer margins on one side and the folded middle at the other,” explained Dudchenko, who also is co-director of DNA Zoo. “And then in other species, each chromosome is crumpled into a little ball.”
“So we had a puzzle,” said Dr. Erez Lieberman Aiden, an associate professor of molecular and human genetics and Emeritus McNair Scholar at Baylor, co-director of the DNA Zoo and senior author on the new study. “The data implied that over the course of evolution, species can switch back and forth from one type to the other. We wondered: What is the controlling mechanism? Might it be possible to change one type of nucleus into another in the lab?” Aiden also is director of the Center for Genome Architecture and a senior investigator at CTBP.

Condensin II influences chromosome arrangement

Meanwhile, an independent team in the Netherlands had discovered something unexpected. “I was doing experiments on a protein called condensin II, which we knew plays a role in how cells divide,” said Claire Hoencamp, co-first author of the study and a member of the laboratory of Dr. Benjamin Rowland at the Netherlands Cancer Institute. “But we observed the strangest thing: When we mutated the protein in human cells, the chromosomes would totally rearrange. It was baffling!”
The two teams met at a conference in the Austrian mountains, where Rowland presented his lab’s latest work. They soon realized that Hoencamp had hit on a way to convert human cells from one nuclear type to another.
When we looked at the genomes being studied at the DNA Zoo, we discovered that evolution had already done our experiment many, many times! When mutations in a species break condensin II, they usually flip the whole architecture of the nucleus,” said Rowland, senior author on the study.
“It’s always a little disappointing to get scooped on an experiment, but evolution had a very long head start.”

The team decided to work together to confirm condensin II’s role. But then the COVID-19 pandemic struck, and much of the world shut down.
“Without access to our laboratories, we were left with only one way to establish what condensin II was doing,” Hoencamp said. “We needed to create a computer program that could simulate the effects of condensin II on the chain of hundreds of millions of genetic letters that comprise each human chromosome.”

The team turned to Dr. José Onuchic, the Harry C. and Olga K. Wiess Chair of Physics at Rice. “Our simulations showed that by destroying condensin II, you could make a human nucleus reorganize to resemble a fly nucleus,” said Onuchic, co-director of CTBP, which includes collaborators at Rice, Baylor, Northeastern University and other institutions in Houston and Boston.
The simulations were performed by a team within Onuchic’s lab at CTBP, led by postdoctoral fellow and co-first author Dr. Sumitabha Brahmachari, working with Dr. Vinicius Contessoto, a former postdoc at CTBP, and Dr. Michele Di Pierro, a CTBP senior investigator and currently an assistant professor at Northeastern University.
“We began with an incredibly broad survey of 2 billion years of nuclear evolution,” Brahmachari said.

We found that much boils down to one simple mechanism, that we can simulate as well as recapitulate, on our own, in a test tube.”
“It’s an exciting step on the road to a new kind of genome engineering — in 3D!”
Work at Rice, Baylor and Northeastern was supported by the National Science Foundation (NSF), the Welch Institute, the National Institutes of Health, the NSF-supported Behavioral Plasticity Research Institute, IBM, the Pawsey Supercomputing Center and Illumina Inc.
Read the paper’s abstract here.