SPOTlight can identify individual cells among hundreds of thousands
Finding and isolating single live cells with unique profiles from heterogenous populations is now possible with SPOTlight. This novel platform was developed by Rice University graduate student Jihwan (James) Lee and Dr. François St-Pierre, an assistant professor of neuroscience at Baylor College of Medicine and an adjunct assistant professor of electrical and computer at Rice.

In the process, the researchers found a more robust fluorescent protein. Both the platform and the protein could be highly useful to synthetic biologists and biomedical researchers. They often need to single out cells with specific visual characteristics like shape or activity determined by their genetic or epigenetic makeup or their developmental history.
Lee and his colleagues dubbed their platform SPOTlight, short for Single-cell Phenotypic Observation and Tagging with Light. It addresses the limitations of existing sorting techniques to isolate single live cells with unique profiles from heterogenous populations. They then leveraged the method for protein engineering to develop the most photostable yellow fluorescent protein reported to date.
“We basically developed a platform that allows one to screen for spatial and temporal properties of individual cells,” said Lee, the first author and a student in Rice’s Systems, Synthetic and Physical Biology program working in St-Pierre’s Baylor lab.
How SPOTlight works
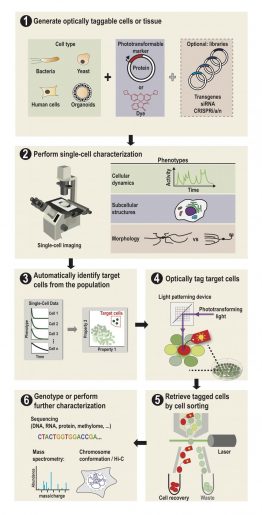
“This is done by first observing the cells under a microscope,” Lee said. “The cells express a special protein so that shining a spot of light on desired cells make them go red. We can then easily separate red cells from the rest using a common device called a flow cytometer.”
That “special” photoactivatable fluorescent protein irreversibly transitions from dark to bright after being zapped by violet light. Photoactivatable dyes can also be used instead of proteins. In effect, cells are left with a long-lasting tag.To only tag cells of interest, the team used a digital micromirror device, an array of tiny motor-driven mirrors also used in digital projectors, to give it the ability to light up single cells.
“These micromirrors rotate and turn to define a region of your sample, down to single cells,” Lee said. “This is all automated. There’s a motorized microscope stage that moves the cells on an imaging plate around a predefined zone, and the DMD will shine light only on a particular cell.”
Through SPOTlight, a researcher can observe a population of hundreds of thousands of human or yeast cells over time to find those with desirable cellular dynamics, subcellular structures or shapes.
Custom software can then be used to identify all cells with the desired profile, and instruct the light source and the DMD to photoactivate them with violet light.
“Then we use a flow cytometer or cell-sorting machine that can detect and recover the cells we tagged while throwing away the rest,” Lee said. “After we’ve recovered our cells of interest, we can send them for sequencing or conduct further studies.”
Lee said the prototype tags individual cells in 45 seconds to a minute. “That depends on the power of the light,” he said. “With a stronger light source, we should be able to do this even faster, maybe down to a few seconds per cell.”
Improving the fluorescent tag
To demonstrate the utility of SPOTlight, Lee and his colleagues used it to screen 3 million mutant cells expressing a library of fluorescent proteins, ultimately identifying and refining a yellow fluorescent protein they call mGold.
“It’s a variant of an existing fluorescent probe called mVenus,” Lee said. “The problem with mVenus is that it photobleaches very fast. It becomes dimmer and dimmer as you keep shining light on it. If you’re monitoring cells expressing mVenus for a long time, there comes a time where the fluorescent protein is no longer detectable. So we decided to screen for mVenus mutants with better fluorescent stability.”
He said researchers typically engineer fluorescent proteins by shining light on bacterial colonies expressing the proteins to see which one is brightest. With SPOTlight, “we can screen for brightness and photostability at the same time,” Lee said. “This isn’t something people commonly did, but biology isn’t static. It’s moving in time and space, so it’s important to have these temporal properties as well.
“Compared with commonly used yellow fluorescent proteins, mGold was four to five times more stable,” he said.
“Important developmental events and behaviors require monitoring for many minutes, hours or days and it’s frustrating when the probes we use to image these processes go dark before we’ve been able to capture the whole story,” said St-Pierre, McNair Scholar at Baylor.
It’s like having a power outage in the middle of watching a good movie,” he said. “Building on our work with mGold, we now want to use SPOTlight to develop probes that will enable us to watch full movies.”
“Similarly, SPOTlight can enable synthetic biologists to engineer new proteins, nucleic acids or cells,” St-Pierre said. “More broadly, this method can help any researcher seeking to unravel the genetic or epigenetic determinants of an interesting cellular phenotype, including such clinically relevant properties as resistance to disease or treatment.”
The team reported their results in Science Advances.
Co-authors of the paper are Rice graduate students Zhuohe Liu and Xiaoyu Lu, undergraduate John Ahrens and alumnus Peter Suzuki, and research assistant Shujuan Lai and alumna Sihui Guan of Baylor.
This research was supported by Baylor College of Medicine, the McNair Medical Foundation, the Welch Foundation, the Klingenstein-Simons Foundation, the Cancer Prevention and Research Institute of Texas, the National Science Foundation and the National Institutes of Health.
Read more here.


Wow, this is amazing