Image of the Month: Locating molecular players in Batten disease
Mutations in either protein CLN6 or CLN8 result in two forms of Batten disease with remarkably similar clinical features. It turns out, as recently shown by the laboratory of Dr. Marco Sardiello, that both proteins work together to equip lysosomes, the waste-disposal hubs of the cell, of the much needed enzymes that process cellular waste.
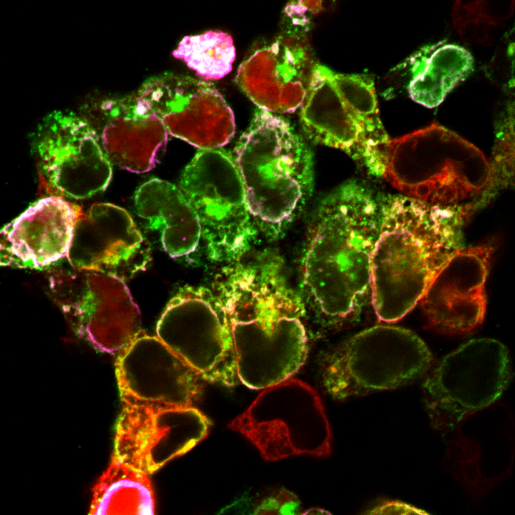
Batten disease is a family of 13 rare, genetically distinct conditions. Collectively, they are the most prevalent cause of neurodegenerative disease in children, affecting 1 in 12,500 live births in the U.S. One of the Batten disease genes is CLN6. How mutations in this gene lead to the disease has been a mystery, but a study led by researchers at Baylor College of Medicine and published in the Journal of Clinical Investigation reveals how defective CLN6 can result in Batten disease.
“People with Batten disease have problems with their cells’ ability to clear cellular waste, which then accumulates to toxic levels,” said first author Dr. Lakshya Bajaj, who was working on this project while a doctorate student in the Sardiello lab at Baylor. Bajaj is currently a post-doctoral associate at Harvard Medical School.

In cells, lysosomes process cellular waste. They are sacs containing enzymes, a type of proteins that break down waste products into its constituent components that the cell can recycle or discard. In Batten disease caused by mutations in CLN6, the lysosomes do not process waste effectively for unknown reasons. This results in waste accumulation. Batten disease is a type of lysosomal storage disorder. Although all types of cells can be affected by defects in lysosomal waste management, brain cells, neurons, are particularly susceptible.
Waste accumulation in neurons perturbs many cellular processes and eventually results in cell death. This leads to the progressive degeneration of motor, physical and intellectual abilities observed in Batten disease patients,” Bajaj said.
CLN6: another piece of the Batten disease puzzle
The connection of CLN6 with Batten disease was a bit of a mystery. This protein is not found in lysosomes, but in the endoplasmic reticulum, a structure inside cells where proteins, including lysosomal enzymes, are made. The endoplasmic reticulum is separate from the lysosomes. So, how do defects in a protein located outside of the lysosomes interfere with lysosomal function?
The Sardiello lab had previously solved a similar mystery involving CLN8, another protein located in the endoplasmic reticulum and whose mutations also cause a type of Batten disease.
“We showed that CNL8 assists on the exit of lysosomal enzymes from the endoplasmic reticulum en route to the lysosomes. When CLN8 is defective, the transport of enzymes from their place of synthesis to the final destination is deficient and the lysosomes end up having fewer enzymes to work with,” said Sardiello, associate professor of molecular and human genetics at Baylor and corresponding author of this work.

CLN6 and CLN8 work together
The clinical manifestations of Batten disease caused by CLN8 mutations and those of Batten disease due to defective CLN6 are remarkably similar. This and other evidence led the researchers to suspect that CLN6 and CLN8 might be working together.
Their investigations revealed that CLN6 and CLN8 do interact with each other forming a molecular complex that collects lysosomal enzymes at the endoplasmic reticulum and mediates their trafficking towards the lysosomes.
“We propose that CLN8 and CLN6 together herd the enzymes into a hub, a sort of ‘bus stop.’ Then, CLN8 escorts the enzymes on the bus en route to the lysosomes, while CLN6 remains at the bus stop. CLN8 returns to the bus stop after delivering the enzymes, and they repeat the process,” Bajaj said. “When CLN6 is defective, the enzymes are not effectively herded into the bus stop and fewer are transported to the lysosomes.”
The researchers are interested in finding whether other factors are involved in transporting enzymes to the lysosomes. For instance, whether there are other ‘bus conductors or herders’ of lysosomal enzymes involved that, if defective, may also contribute to Batten disease.
Other contributors to this work include Jaiprakash Sharma, Alberto di Ronza, Pengcheng Zhang, Aiden Eblimit, Rituraj Pal, Dany Roman, John R. Collette, Clarissa Booth, Kevin T. Chang, Richard N. Sifers, Sung Y. Jung, Jill M. Weimer, Rui Chen and Randy W. Schekman. The authors are affiliated with one or more of the following institutions: Baylor College of Medicine; Texas Children’s Hospital; University of California, Berkeley; Sanford Research, Sioux Falls, South Dakota; and Sanford School of Medicine at the University of South Dakota.
This work was supported by NIH grants NS079618 and GM127492 and grants from the Gwenyth Gray Foundation, Beyond Batten Disease Foundation and NCL-Stiftung. This project was supported in part by IDDRC grant number 1U54 HD083092 from the Eunice Kennedy Shriver National Institute of Child Health and Human Development, the Integrated Microscopy Core and the Proteomics Core at Baylor College of Medicine with funding from NIH (DK56338, and CA125123), CPRIT (RP150578, RP170719), the Dan L Duncan Comprehensive Cancer Center and the John S. Dunn Gulf Coast Consortium for Chemical Genomics.