Solving the puzzle of Mitchell disease
When a patient with puzzling neurological symptoms enrolled in the Undiagnosed Diseases Network, researchers led by Dr. Hugo J. Bellen were set on solving the mystery. The patient presented with an unidentified late-onset neurodegenerative disorder. The team named this new syndrome “Mitchell disease” in reference to the first patient to be diagnosed with this disorder and looked to identify its genetic basis.
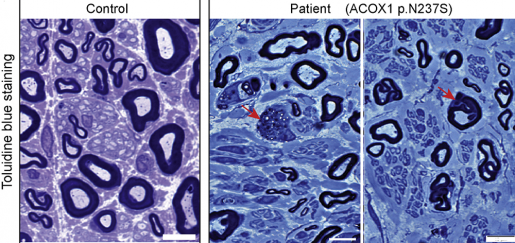
“On comparing the patient’s and his parents’ DNA, the team identified a mutation in the patient that resulted in a single amino acid substitution (N237S) in the ACOX1 protein. This change was seen only in the patient and was not present in either of his parents’ DNA, indicating that the patient had a de novo, or new, mutation on this gene,’ said Bellen, professor at Baylor College of Medicine and investigator at the Jan and Dan Duncan Neurological Research Institute at Texas Children’s Hospital and also a Howard Hughes Medical Institute investigator. “With the help of the online gene-matching tool GeneMatcher, we found two more patients who had the same new mutation in the ACOX1 gene.”
All three patients, who ranged from 3 to 12 years old at the time of disease onset, had remarkably similar clinical features, including degeneration of peripheral nerves that caused a progressive loss of mobility and hearing. The three individuals had identical gene variants, a clear indication that ACOX1 dysfunction likely was the cause of the symptoms.
A medical mystery
The finding that an ACOX1 mutation was linked to Mitchell disease initially baffled the researchers. The only known ACOX1-related disorder described in the medical literature at that time presented earlier in infancy with seizures, severe cognitive decline, neuro-inflammation and accumulation of very-long-chain-fatty acids in plasma and, more importantly, was caused by the lack of the ACOX1 protein – none of which was true for these three patients.
“The brain has large amounts of lipids, which are critical for the proper functioning of the nervous system. Abnormal breakdown of lipids in the brain and peripheral nervous system is associated with several neurodegenerative diseases,” Bellen said.

The gene ACOX1 is involved in lipid breakdown. It produces an enzyme called Acyl-CoA oxidase 1 that initiates a series of reactions that break down very-long-chain-fatty acids in small intracellular organelles called peroxisomes.
Fruit flies help solve the medical mystery
To resolve this conundrum, the Bellen team turned to fruit flies. The first surprising discovery made by the lead author, Hyunglok Chung, was that the ACOX1 protein is abundant and critical for the maintenance of glia, cells that support neurons. This uncovered a previously unknown role of peroxisomes in glial cells and paved the way for further experiments.
To understand how ACOX1 variants affect the function of glia, they generated two mutant fly lines, the first one lacked both the copies of ACOX1 gene and the second, carried the substitution mutation (N237S) found in one of the ACOX1 genes in the Mitchell disease patients.
“Flies lacking ACOX1 mimicked the symptoms of ACOX1 deficiency in humans, including elevated levels of very-long-chain-fatty acids along with dramatic loss of glia and neurons and progressively impaired neuronal function. When we reduced the synthesis of very-long-chain-fatty acids in these flies by administering the drug bezafibrate, we observed significant improvement in lifespan, vision, motor coordination and neuronal function, implicating elevated levels of these lipids and their excessive accumulation in glia as an important contributor,” said Chung, postdoctoral fellow in the Bellen lab.
It is remarkable how well bezafibrate suppressed the symptoms of ACOX1 deficiency, suggesting a new therapeutic avenue for patients with this condition,” Bellen said.
In contrast to the loss of ACOX1, the introduction of the single amino acid substitution (N237S) in ACOX1 gene resulted in a hyperactive ACOX1 protein. Typically, breakdown of very-long-chain-fatty acids by the enzymatic action of ACOX1 produces small amounts of highly reactive oxygen species, but glial cells quickly neutralize them. However, in Mitchell’s disease, hyperactive ACOX1 produces copious amounts of toxic reactive oxygen species, leading to the destruction of glia and their neighboring neurons.
The harmful effects due to hyperactive ACOX1 were potently reversed with the antioxidant N-acetyl cysteine amide (NACA). However, NACA did not suppress the lethality or toxic effects in flies that lacked ACOX1, a clear indication that the two diseases act via entirely different pathways and would need to be treated with two distinct therapeutic strategies.
“This study is a prime example of how combining UDN’s unique team science approach with power of fruit fly genetics is facilitating rapid and phenomenal progress in rare diseases research. We take on cases of patients with conditions never described before, uncover new diseases and find definitive molecular diagnosis for them. We make significant progress in unraveling the causes of these novel diseases and rapidly identify and test promising new treatment options,” Bellen said. “We have successfully identified more than 25 disease-causing genes within the past three years – a task that typically takes many years.”
The study appears in the journal Neuron.
Other contributors to this study include Michael F. Wangler, Paul C. Marcogliese, Juyeon Jo, Thomas A. Ravenscroft,, Zhongyuan Zuo, Lita Duraine, Sina Sadeghzadeh, David Li-Kroeger, Robert E. Schmidt, Alan Pestronk, Jill A. Rosenfeld, Lindsay Burrage, Mitchell J. Herndon, Shan Chen, Undiagnosed Diseases Network, Amelle Shillington, Marissa Vawter-Lee, Robert Hopkin, Jackeline Rodriguez-Smith Michael Henrickson, Brendan Lee, Ann B. Moser, Richard O. Jones, Paul Watkins, Taekyeong Yoo, Soe Mar, Murim Choi, Robert C. Bucelli, Shinya Yamamoto, Hyun Kyoung Lee, Carlos E. Prada, Jong-Hee Chae, and Tiphanie P. Vogel.
The authors are affiliated with one or more of the following organizations: Baylor College of Medicine, Texas Children’s Hospital, Washington University in St. Louis, Cincinnati Children’s Hospital Medical Center, Johns Hopkins School of Medicine, and Seoul National University College of Medicine.
This work is funded by NMSS via RG1508-08406, NIH Common Fund, through the Office of Strategic Coordination/Office of the NIH Director under Award Number(s) U01HG007709, U54NS093793 and by ORIP via R24OD022005, BCM IDDRC (supported by NICHD U54HD083092), Howard Hughes Medical Institute and Cullen Foundation.