Not in sync: single cells and genes show unanticipated response to estrogen
When experimental findings are not compatible with the textbook view, scientists know they have quite a challenge in their hands. Dr. Fabio Stossi, Dr. Michael A. Mancini at Baylor College of Medicine and their colleagues faced this situation when they were investigating how cells modulate their responses to estrogen at single-cell and -gene level.
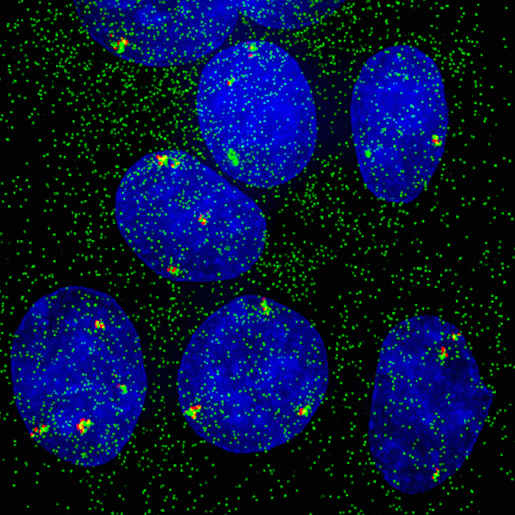

“Estrogen is a type of steroid hormone that modulates a large number of biological functions, both in males and females, by regulating the activity of hundreds of genes per cell,” said Stossi, associate professor of molecular and cellular biology and technical director of the Integrated Microscopy Core at Baylor.
A great deal is known about how estrogen triggers its effects. It binds to a nuclear transcription factor (estrogen receptor, or ER), which in turn interacts with specific DNA sequences facilitating the recruitment of coregulators that participate in the regulation of gene expression.
It was assumed that this process would likely happen simultaneously in all the ER-containing cells in a population that was stimulated with estrogen, but little was known of how actual single cells or individual alleles – copies of the same gene – responded. Which is why researchers did not anticipate their findings.
Looking at individual cells and alleles

“In the current study, we worked with human breast cancer cell lines grown in the lab. Using both molecular and imaging analyses, we determined, at single-cell and -allele levels, the expression of two well-characterized genes, GREB1 and MYC, whose activity is regulated by estrogen,” said Mancini, professor of molecular and cellular biology, and pharmacology and chemical biology at Baylor. Mancini also is the academic director of the long-running Integrated Microscopy Core (IMC) at Baylor and director of the recently-formed GCC Center for Advanced Microscopy and Image Informatics (CAMII), a CPRIT-funded resource across Baylor and the Texas A&M Institute for Bioscience and Technology.
The researchers incubated the cells in the lab and treated them with estrogen. Then, they looked at the expression of GREB1 and MYC genes in individual cells, and at the expression of individual alleles in each cell. As expected, they found that estrogen activated GREB1 and MYC genes quickly, within 15 minutes, but there was an unexpected and marked asynchronous response to hormone simulation at both individual cell and allele levels.
“Our analyses showed that gene activation of cells in a population appeared more random than we expected. In single cells, the response of each copy of the gene was independent from that of its neighboring cells. In some cells, no alleles of the genes were active, whereas cells next to them would have some or all their gene copies active,” Stossi said.
Nevertheless, the researchers explained, although these findings had not been described before in estrogen receptor biology, they were not a complete surprise.
Studies in genetically identical bacteria have shown that, when subjected to the same treatment, not all the bacteria respond the same way. This is called phenotypical heterogeneity.
“We have been interrogating mechanisms of action of ER via state-of-the-art imaging/analysis since the 90’s, with continual improvements in our resources as they are developed or when they come to market,” Mancini said.
In this study we have uncovered novel characteristics of estrogen action in mammalian cells for the first time.”
“Having phenotypical heterogeneity confers an important adaptation strategy to cell populations, whether they are cancerous or normal. If all the cells in a population responded the same way to a harmful stimulus, for instance, by stopping an essential function, then they would not have the capability of surviving. But responding differently may allow some cells to survive,” Mancini said.
A unique opportunity to explore what causes an asynchronous response
The next experiments intended to determine what caused asynchronous estrogen-triggered gene activation. Since the imaging and image analysis routines that were developed and used were amenable to the fully automated, high throughput imaging/analysis platforms within the IMC and CAMII, Stossi, Mancini and their colleagues had a unique opportunity to explore this question.
First, the researchers hypothesized that cells responded differently to estrogen because the number of estrogen receptors per cell varied. They were surprised to find out that the number of the estrogen receptors expressed in cells was not strictly dictating whether a cell was going to activate the target genes. Then, the researchers investigated whether the cellular response to estrogen depended on the activation status of the estrogen receptor using a patient-linked, constitutively-active receptor, but again, they found no correlation.
Next, the researchers explored the possibility that estrogen receptor coregulators were involved in modulating the allele-by-allele response to estrogen. Utilizing the automated high throughput resources of the IMC/CAMII, they tested a collection of small molecule epigenetic inhibitors and identified one, called MS049, an inhibitor of two protein arginine methyltransferases, that markedly increased the expected number of active alleles per cell under estrogen stimulation, in a gene-specific manner.
“For the first time we were able to alter the nature of the estrogenic response at the allele level, indicating that there are pathways that serve as rheostats to maintain variability of response to a stimulus, thus preventing maximal and uniform behavior in a population of cells,” Stossi said. “These findings suggested that modifying the activity of coregulators can tweak the variation of allele-by-allele hormonal responses in a gene-specific manner.”
The study provides novel insights into the complex nature of the regulation of gene expression in mammalian cells, suggesting that mechanisms do indeed exist to modulate hormone receptor responses at the single-cell and -allele level.
Interested in utilizing in your research the high throughput single cell imaging resources (IMC/CAMII) described here? Contact Dr. Mancini (mancini@bcm.edu).
Read all the details about this study in the journal Nucleic Acids Research.
Other contributors to this work include Radhika Dandekar, Maureen G. Mancini, Guowei Gu, Suzanne AW Fuqua, Agostina Nardone, Carmine De Angelis, Xiaoyong Fu, Rachel Schiff, Mark T Bedford, Wei Xu, Hans E Johansson and Clifford C Stephan. The authors are affiliated with one or more of the following institutions: Baylor College of Medicine; The University of Texas MD Anderson Cancer Center, Smithville; University of Wisconsin School of Medicine and Public Health, Madison; LGC Biosearch Technologies; Texas A&M University, Houston and the Gulf Coast Consortia Center for Advanced Microscopy and Image Informatics (CAMII), Houston.
Support for this study was provided by NIH/NIEHS Superfund grant (P42ES027704); CPRIT-funded GCC Center for Advanced Microscopy and Image Informatics (RP170719); NIH grants (DK062248 and R01-CA72038), CPRIT (RP120732), and Breast Cancer Research Foundation. Additional support was provided by NIH Breast Cancer Specialized Programs of Research Excellence grants (P50CA058183, P50CA186784), Susan G. Komen for the Cure Foundation Promise Grant (PG12221410), the Integrated Microscopy Core at Baylor College of Medicine with funding from NIH (DK56338, and CA125123), CPRIT (RP150578), the Dan L. Duncan Comprehensive Cancer Center and the John S. Dunn Gulf Coast Consortium for Chemical Genomics. The Gulf Coast Consortium for Chemical Genomics high throughput screening center is funded by CPRIT (RP110532, RP150578).



