E. coli shows the way to discover cell-made protein carcinogens
When searching for carcinogens, people look at the environment. External factors such as tobacco smoke and sunlight can damage DNA, triggering mutations that can cause cancer. But most DNA damage seems to result from events that occur within cells and is mediated by cellular components, including proteins. Despite the importance of these events, they have not been studied extensively.
“In this study, we discovered a new major class of cancer-promoting genes, by showing that many normal proteins made by our cells can act like carcinogens — they damage DNA and cause mutations,” said Dr. Susan M. Rosenberg, Ben F. Love Chair in Cancer Research and professor of molecular and human genetics, of molecular virology and microbiology and of biochemistry and molecular biology at Baylor. “Cancer is a disease of mutations. A normal cell that has accumulated several mutations in particular genes becomes likely to turn into a cancer cell.”
In this project, Rosenberg and her colleagues set out to uncover proteins that, when overproduced by the cell, cause damage to DNA in ways that can lead to cancer. Overproduction of one or another protein is a frequent cellular event, Rosenberg explained.

To uncover these DNA “damage-up” proteins, the researchers took an unconventional approach. They searched for proteins that promote DNA damage in human cells by looking at proteins that, when overproduced, would cause DNA damage in the bacterium E. coli.
“Although bacteria and people are different, their basic biological processes are similar, so with this approach we thought we might find common mechanisms of DNA damage that could be relevant to cancer,” said Rosenberg, who also is leader of the Cancer Evolvability Program at the Dan L Duncan Comprehensive Cancer Center at Baylor.
E. coli reveals novel candidates for cancer-promoting proteins
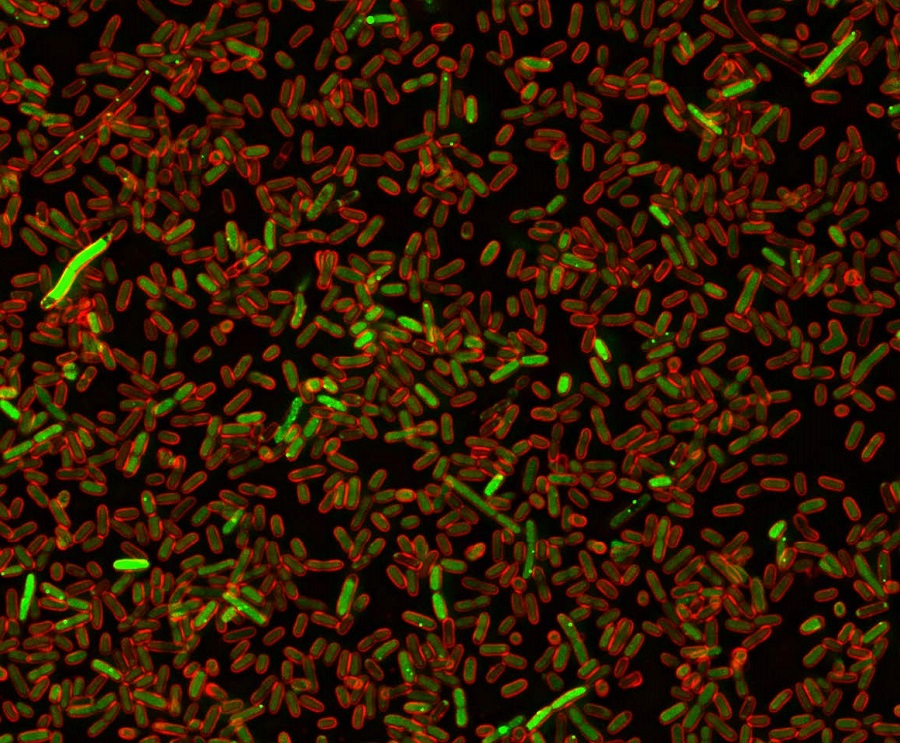
The researchers genetically modified bacteria so they would fluoresce red when DNA was damaged. Then, they overexpressed each of the 4,000 genes present in E coli individually and determined which ones made bacteria glow red.
“We uncovered an extensive and varied network of proteins that can lead to DNA damage,” Rosenberg said. “Some of these proteins are, as expected, involved in DNA processing or repair, but, surprisingly, most are not directly connected to DNA. For instance, some of the DNA damage-up proteins participate in the transport of molecules across the cell membrane.”
Further investigations uncovered 284 human protein relatives of the DNA “damage-up” proteins the researchers had found in bacteria. These human proteins are linked to cancer more often than random sets of proteins. Also, when overproduced in human cells in the lab, half of the proteins triggered DNA damage and mutation.

“E. coli can help to identify DNA damage-up proteins and mechanisms of action in human cells quickly and inexpensively,” said Dr. Christophe Herman, professor of molecular and human genetics and molecular virology and microbiology at Baylor College of Medicine and member of the Dan L Duncan Comprehensive Cancer Center.

“I think it is extraordinary to identify so many ways DNA can be damaged. This study is opening up new avenues for discoveries of novel mechanisms that protect our genomes and how their dysfunction can alter the integrity of our DNA and cause cancer,” said Dr. Kyle M. Miller, associate professor of molecular biosciences at the University of Texas at Austin and member of the Dan L Duncan Comprehensive Cancer Center at Baylor.
“Our work has significant implications both in basic biological fields and in clinical research,” Rosenberg said.
We provide a previously unknown understanding of the diverse cellular mechanisms that can generate DNA damage leading to cancer.”
“In the future, this finding may lead to new ways to identify people who are likely to develop cancer so that strategies to prevent it, slow it down or catch it early can be used,” Rosenberg said.
Would like to know more about this work? Find it in the journal Cell.
The study’s two co-first authors were students earning their doctorates: Dr. Jun Xia at Baylor College of Medicine and Dr. Liya Chiu at the University of Texas at Austin.
This out-of-the-box approach was made possible by funding from two sources aimed at supporting high-risk strategies that, if successful, would have high impact: a National Institutes of Health Director’s Pioneer Award and a gift from the W.M. Keck Foundation, among many other grants to the 16-lab team.
For a complete list of all the contributors and their affiliations and the financial support of this study, follow this link.