The 2018 Michael E. DeBakey M.D. Awards for Excellence in Research
Once again, the 2018 DeBakey Research Awards ceremony fulfilled expectations in all attendees. Not only was the event an opportunity to honor the awardees for their individual outstanding published scientific contributions to basic biomedical research over the past three years, it also was the occasion to learn about the focus of their research and how it has contributed to advance their own fields of study.
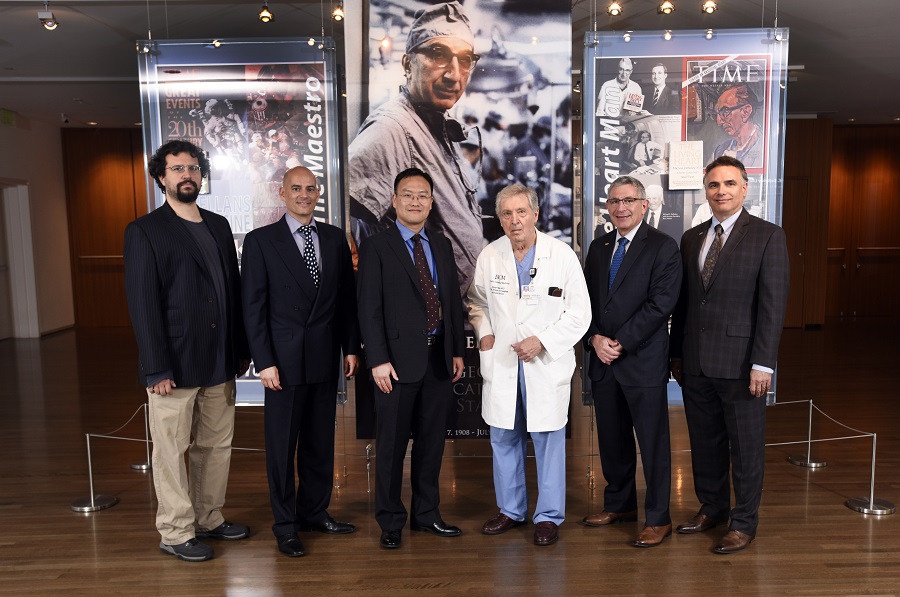

Here are the 2018 awardees, the essence of their corresponding scientific contributions for which they have been honored and a bit of an insight into their talks during the ceremony.
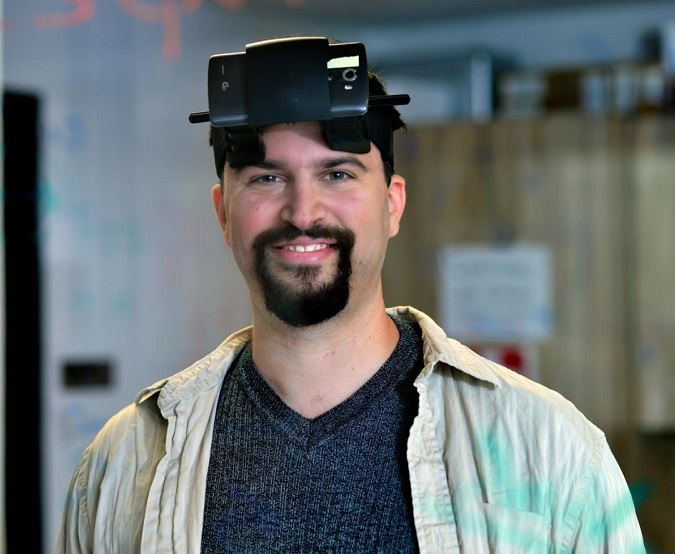

Dr. Erez Lieberman Aiden is assistant professor of molecular and human genetics and a McNair Scholar. His lab is revolutionizing the study of how the human genome, at more than 2 meters long, folds up to fit inside the cell nucleus, and how this process governs gene regulation. His recent work builds on the Hi-C method, which he invented in graduate school.
During his talk, he brought the audience closer to the thinking behind his approach to understanding how DNA folds by comparing it to how Facebook networks help establish associations between people using the example of Homer Simpson’s Facebook connections. In the computer lab, by coupling DNA to DNA proximity ligation and high-throughput sequencing, Hi-C made it possible to solve the problem of how genomes fold. His lab has made a series of transformative discoveries: the compartmentalization of the human genome inside the nucleus; developing the first map of loops in the human genome and discovering how their positions are encoded; the demonstration that these loops form by extrusion, which has enormous consequences for the enzymology of DNA and which is now almost universally accepted; the ability to perform 3-D surgery on a genome by manipulating this code to control the extrusion process; and a method for assembling new, Human Genome Project-quality genomes for $1,000.
Dr. Aiden’s impact is easily quantified. His lab’s paper on mapping loops genome-wide for the first time has been cited more than 1,000 times, the most for any research article in Cell since 2014, and the 3-D genome browser it introduced has been used more than 100 million times. In the last year alone, the Aiden lab has published two papers in Cell, one of which was featured on the cover, and one in Science.

Dr. Karl-Dimiter Bissig is an associate professor of molecular and cellular biology and a member of the Center for Cell and Gene Therapy and the Stem Cells and Regenerative Medicine Center. His research focuses on human liver disease and human disease modeling. His three most recent works were published in Nature Communications.
He told the audience that Leonardo DaVinci has inspired his career. He admires Da Vinci’s courage to explore and accept failure as a teacher. In his first paper, Bissig introduced the first xenograft model for metabolic liver disease. This mouse model more closely resembles human disease modeling, offering the ability to validate human experimental therapies. These new models are of particular relevance to macromolecular therapies, which are sequence or epitope specific, and cannot be validated in experimental animal models. This process supports the rapid translation of therapy from lab to clinical trials. His work was also featured in “research highlights” in Nature Reviews Endocrinology. His follow-up study demonstrated for the first time human-specific drug metabolism in an animal model. He and his team were able to functionally inactivate murine cytochrome metabolism in humanized mice. His results will likely influence drug development and toxicity in a broad sense since most drugs are metabolized through the liver.
In his third paper, he and his colleagues developed metabolic pathway reprogramming uses CRISPR/Cas9 genome editing technology to inhibit an enzymatic pathway rather than to edit a disease-causing gene directly. This publication rescued a lethal phenotype using this technology, and in contrast to common small molecule drugs, required only a single intervention to rescue 100 percent of treated animals.

Dr. Atul Chopra is an assistant professor in the Department of Molecular and Human Genetics and the Department of Molecular and Cellular Biology and is a Caroline Wiess Law Scholar at Baylor.
Dr. Chopra could not attend the ceremony, but his colleague and research collaborator Dr. Yong Xu received the honor in his name and engaged the audience with a presentation about Dr. Chopra’s work. Chopra’s work focuses on metabolism and energy homeostasis. He studies how energy enters the body and how it is used once it’s in the body, which usually involves an interplay between several organs and thus requires a whole-body experimental approach. Chopra also works as a medical geneticist, focusing on patients with specific genetic problems related to consumption of energy, processing of energy and loss of energy. The problems usually manifest as a very high weight or very low weight. Since he was a medical resident at Baylor, he has been working to understand Neonatal Progeroid Syndrome, a condition characterized by energy balance problems resulting in extreme thinness and propensity for hypoglycemia. His work led to the discovery of a new glucogenic and orexigenic hormone, asprosin. This hormone modulates hepatic glucose production and appetite stimulation using spatiotemporally distinct mechanisms when dietary fuel is unavailable. Chopra’s first paper on the glucogenic effect of asprosin in the liver was published in Cell and the second, which characterized the orexigenic effect of asprosin in the hypothalamus, was published in Nature Medicine.
This discovery of asprosin represents a new direction in the understanding of Marfan syndrome and the study of fibrillin. Asprosin has the potential for beneficial therapeutic impact of activity blockade via monoclonal antibody. In addition, antibodies targeting asprosin are showing considerable promise by acutely lowering blood glucose and appetite, leading to improvements in type II diabetes and obesity, respectively, over time.

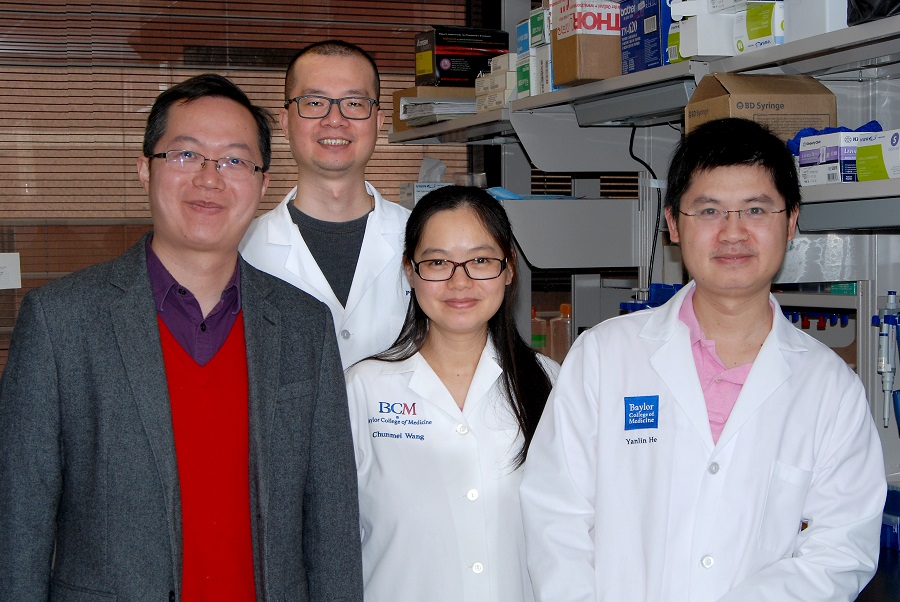
Dr. Meng Wang, is associate professor of molecular and human genetics and the Fyfe Endowed Chair in the Huffington Center on Aging. She has developed a successful innovative multidisciplinary research program that has produced original contributions to the fields of aging biology, lipid metabolism and reproductive biology.
Dr. Wang could not attend the event, but had prepared a video of her talk in which she told us how her laboratory uncovered the first lysosome-to-nucleus retrograde lipid messenger pathway, provided in-depth biochemical mechanisms for its action, and demonstrated its novel roles in regulating longevity. (Science, 2015). Her work published in Nature Cell Biology and Cell also provided the first evidence for a novel communication mode between bacteria and their eukaryotic mitochondria, deciphered bacteria-derived metabolites mediating this ancient dialogue and their signaling mechanisms, and determined their vital effects on host’s lipid metabolism and longevity. These studies have direct translational impact as they identify new nutrient-based approaches to increasing health span. In the area of reproductive fitness, she conducted the first genome-wide RNAi screen for gene inactivation that induces reproductive longevity, and identified a list of new genes associated with endocrine control of reproductive aging.
Given the increasing trend of delayed childbearing in our current society, her discoveries are timely and lay critical foundations for improving reproductive health in humans. Her groundbreaking contributions to technological innovation now allow for sensitive and specific trace of in vivo dynamics of small metabolites in living cells and organisms. These new technological advances have transformed how we investigate these bioactive small molecules and have revealed previously unknown physiological activities.

Dr. Xiang Zhang is an associate professor of molecular and cellular biology, a McNair Scholar and a member of the Dan L Duncan Comprehensive Cancer Center. He has focused the efforts of his lab in elucidating biological mechanisms and therapeutic strategies of metastasis, the major threat to the lives of solid cancer patients.
Through his presentation he connected for the audience the different approaches his lab has taken to improve the way we fight breast cancer. These efforts resulted in the development of a set of unique techniques and models to study the interaction between microscopic metastases and various stromal components in different organs, particularly in the bone. These investigations have subsequently led to the discovery of osteogenic cells as the major microenvironmental components that promote early-stage of breast cancer-bone colonization, work that was published in Cancer Cell and selected as one of the journal’s eight best papers in 2015. Dr. Zhang is also studying the interactions between cancer cells and various immune cells and has published this work in Nature Cell Biology. Specifically, he has focused on the heterogeneity of the immune environment created by cancers with diverse genetic and epigenetic backgrounds and the establishment of a link between oncogenic mTOR signaling and accumulation of myeloid-derived suppressor cells.
A breakthrough contribution to his field, which was published in a study in Nature, has been the discovery and characterization of a mutually regulatory loop between tumor vasculature and adaptive immunity. These findings profoundly enriched our understanding of tumor-microenvironment interaction and proposed compelling therapeutic strategies.
Interested in knowing about the awardees of previous years? Visit the webpage of the Michael E. DeBakey M.D. Excellence in Research Award.