Researchers make strides identifying genetic causes of rare neurodevelopmental disorders in the Turkish and worldwide populations
Identifying the genetic causes of rare neurodevelopmental disorders can be quite challenging. In a recent study, a global scientific team including researchers from Baylor College of Medicine, worked to find genetic answers for Turkish families.

“It’s very common in clinical practice to see a patient whose characteristics do not match what has been documented in the literature, limiting the physician’s ability to guide clinical care and provide information about which other family members might be at risk,” said Dr. Tadahiro Mitani, a postdoctoral associate in Dr. James R. Lupski’s lab at Baylor College of Medicine.
In the current study, explains Mitani, who is the first author of the work by a team of 50 investigators from around the world, the researchers looked to identify the genetic causes of rare neurodevelopmental disorders in 234 subjects and 20 previously unsolved cases of affected families of the Turkish population.

“To achieve this goal, we integrated improved genome-wide screening technologies, including exome sequencing and whole-genome sequencing, and newly developed computational tools and bioinformatic analyses to improve our ability to identify the genetic underpinnings of rare neurodevelopmental conditions,” said co-corresponding author Dr. Davut Pehlivan, assistant professor of pediatrics – neurology at BCM.
The researchers started this project in 2011 and over the years developed close collaborations with physicians and patients worldwide, as well as with researchers in the fields of genetics, genomics and bioinformatics. The team used GeneMatcher, a freely accessible web-based ‘matchmaking service’ designed to enable connections between clinicians and researchers from around the world who share an interest in the same gene or genes.
Improved diagnosis
The team identified new genes and confirmed genes previously associated with neurodevelopmental disorders.
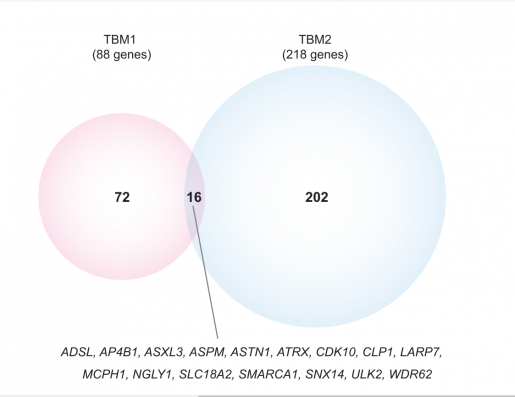
They were able to make a molecular diagnosis in 181 of 254 (71%) of the individuals in this study and in approximately 80% of neurodevelopmental disorders overall. Twenty of the 181 diagnosed individuals had been studied before, but at the time the researchers did not identify a genetic diagnosis.
Our findings confirm that applying newly developed molecular and computational tools on existing data can provide answers to previously undiagnosed families,” Pehlivan said.
A new perspective of complex neurodevelopmental disorders
“Importantly, we also found an explanation for the diagnostic challenge presented by conditions with characteristics that do not match what has been reported in the medical literature,” said Mitani, currently at Jichi Medical University, Tokyo, Japan. “We determined that the accumulation of particular combinations of rare disease-causing gene mutations at multiple genes, a phenomenon called multilocus pathogenic variation, results in complex characteristics unique to each individual.”
The original idea that a single disorder is caused by a mutation in a single gene does not explain the variety of complex neurodevelopmental disorders,” Pehlivan explained.
In multilocus pathogenic variation, one patient may have multiple mutated genes. For instance, one gene mutation may result in muscle disease and a different gene mutation that leads to brain disease, while in another patient one mutation may affect the kidneys and another the brain.
The accumulation of specific combinations of rare multiple mutated genes results in conditions with complex characteristics that are unique to each individual.
Patients may present with neurodevelopmental disorders that share similarities but also have important differences, which need to be taken into consideration when deciding treatment and when evaluating risk for other family members.
“In this study, for the first time we strictly applied a set of criteria to evaluate multilocus pathogenic variation in our patients and found that it was present in 28.9% of the cases in which we established a genetic diagnosis,” Pehlivan said. “Our findings confirm the value of routinely applying these criteria to assess the contribution of multilocus pathogenic variation to rare neurodevelopmental disorders and again revealed why genomic studies are superior to single gene testing.”

“The integrated analyses of the genetic and genomic characteristics of each patient enabled the team to improve their ability to reach a diagnosis in many cases,” said co-author Dr. Zeynep Coban Akdemir, assistant professor at UT Health School of Public Health-Houston. “Most patients with multilocus pathogenic variation are in consanguineous families.”

“With studies such as this one, we seek to tackle the challenge of finding the cause of currently unexplained rare genetic disorders,” said co-author Dr. Jennifer Posey, assistant professor of molecular and human genetics at BCM. Posey also leads the newly launched BCM – GREGoR (Genomic Research to Elucidate the Genetics of Rare) program, a part of the NIH-funded GREGoR Consortium.
The researchers’ comprehensive approach also adds a valuable resource of information to the study of the function of human genes, human biology and molecular mechanisms involved in neurodevelopmental disorders, all of which can lead to improved diagnosis and treatments.
For a complete list of the contributors to this paper, their affiliations and the financial support for the work, see the publication in The American Journal of Human Genetics.