Innovative, faster and more cost-effective technology identifies drug that inhibits SARS-CoV-2 growth in the lab
Much progress has been made on the development of vaccines aiming at triggering human immunity to SARS-CoV-2, the virus that causes COVID-19. In addition, researchers at Baylor College of Medicine are involved in the development of antiviral drugs to treat infected individuals.
Unfortunately, traditional drug development often involves a slow and expensive process that is ill-suited to the public health demands of this global pandemic.
Baylor researchers turned to an innovative, faster and more cost-effective drug discovery technology to screen billions of compounds for a potential treatment against viruses. In doing so, they were able to identify an inhibitor of a viral protein that is needed for propagation of viruses, specifically SARS-CoV-2. That viral protein is called Mpro.
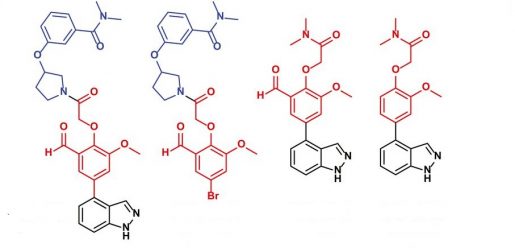
What is Mpro, and why focus on this as a drug target for coronavirus?
“Viruses like SARS-CoV-2 have to reproduce themselves inside of human cells, so if you can find molecules that block key parts of that process then you can prevent the virus from propagating,” said Dr. Damian Young, assistant professor of pharmacology and chemical biology, associate director of the Center for Drug Discovery at Baylor and co-contributing author on this paper.

The method that Young and his colleagues used to screen their compounds is not entirely new and has been mostly applied to treating different forms of cancer. This is the first time that it has been reported to be effectively applied to SARS-CoV-2.
What is unique about this screening process?
The more commonly used method for discovering drugs is called High Throughput Screening, which involves screening at most a million compounds in individual test tubes.
But in this case the researchers used a method called DNA-Encoded Chemistry Technology, which enabled them to screen 4 billion DNA-encoded molecules all in one test tube against a key SARS-CoV-2 protein to find a molecule that may modify how that protein behaves.
“It is a process that allows us to test over 1,000 times more compounds than the traditional process and at a much faster pace. We applied it to finding molecules that hit the viral protein Mpro,” said Young.
How did you screen for the Mpro inhibitor?
Mpro is the SARS-CoV-2 main protease, a key component for the lifecycle of coronaviruses and a therapeutic target that is independent of the current vaccine strategies.

“By blocking Mpro with small molecules, we are blocking SARS-CoV-2 from replicating in cells. These studies are highly encouraging and indicate that this process could be used in the future for emerging viruses, including other coronaviruses,” said Dr. Srinivas Chamakuri, an instructor in pathology and immunology at Baylor and co-first author on this paper.
“The DNA-Encoded Chemistry Technology process involves a simultaneous screen of billions of molecules in which each drug-like molecule is tagged with a DNA barcode,” said Dr. Martin Matzuk, professor and chair of pathology and immunology, director of the Center for Drug Discovery at Baylor and co-contributing author. “The molecules that ‘stick’ to the protein (in this case SARS-CoV-2 Mpro) are identified by sequencing of their attached DNA ‘barcode.’ This is a rapid drug discovery screen, and our study demonstrates the enormous potential to find unique molecules that can inhibit viruses and help in this and in future pandemics.”

The next steps to further test the inhibitor against Mpro will involve animal models and other lab safety tests before it can begin human trials.
Why is this research important when faced with virus mutations?

“We need to find an effective treatment for COVID-19 in addition to the use of vaccines,” said Dr. Timothy Palzkill, professor and chair of pharmacology and chemical biology and co-contributing author. “The rapid rise of variants with mutations that we are observing in SARS-CoV-2 show us the need to be able to do these types of screenings in a faster, more efficient manner.”
“We are very happy to release our research to the public domain. We believe this will be helpful to the scientific community around the world and to those groups working on finding small molecule drugs to fight against SARS-CoV-2 ,” added Chamakuri. “There are multiple people from different specialties that came together at the Center for Drug Discovery at Baylor to support this effort at a time when so many people are suffering.”
Their findings are in the Proceedings of the National Academy of Sciences.
Other authors who contributed in the study include: Shuo Lu, Melek Nihan Ucisik, Kurt M. Bohren, Ying-Chu Chen, Huang-Chi Du, John C. Faver, Ravikumar Jimmidi, Feng Li, Jian-Yuan Li, Pranavanand Nyshadham, Stephen S. Palmer, Jeroen Pollet, Xuan Qin, Shannon E. Ronca, Banumathi Sankaran, Kiran L. Sharma, Zhi Tan, Leroy Versteeg and Zhifeng Yu, all with Baylor College of Medicine or Texas Children’s Hospital. Palzkill also holds the Cullen Trust for Higher Education Academic Chair at Baylor. Matzuk also holds the Stuart A. Wallace Chair and the Robert L. Moody Sr. Chair. Matzuk and Young are members of the Dan L Duncan Comprehensive Cancer Center.
Funding came from the National Cancer Institute (grant P20CA221729), a Core Facility Support Award from the Cancer Prevention and Research Institute of Texas (grant RP160805), the Welch Foundation (grant H-Q-0042), and an internal pilot grant from Baylor College of Medicine.