Proteomics analysis identifies potential drug targets for aggressive human cancers
Historically, researchers have mostly been generating transcriptomic data (the messenger RNA (mRNA) that is translated into protein) from cancer tissues seeking to identify clues about altered cellular pathways that might be driving cancer behavior and could represent novel therapeutic targets. Although this approach has been quite useful, researchers at Baylor College of Medicine think that transcriptomic data alone might not reveal the whole story behind cancer.
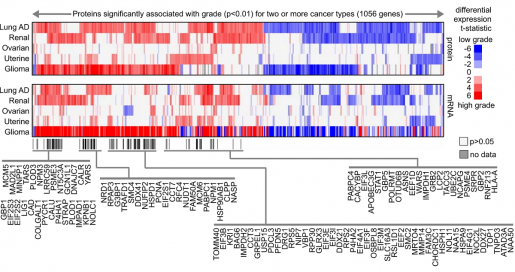
“There are two notable aspects of this study. One is that we explored the proteomic landscape of cancer looking for proteins that were expressed in association with aggressive forms of the disease,” said Dr. Chad Creighton, professor of medicine and co-director of Cancer Bioinformatics at the Dan L Duncan Comprehensive Cancer Center at Baylor. “We analyzed protein data that included tens of thousands of proteins from about 800 tumors including seven different cancer types – breast, colon, lung, renal, ovarian, uterine and pediatric glioma – made available by the Clinical Proteomic Tumor Analysis Consortium (CPTAC) mass-spectrometry-based proteomics datasets.”

Computational analysis for the CPTAC datasets identified proteomic signatures associated with aggressive forms of cancer. These signatures pointed at altered cellular pathways that might be driving aggressive cancer behavior and could represent novel therapeutic targets. Some signatures included cellular pathways of altered metabolism. Each cancer type showed a distinctive proteomic signature for its aggressive form. Interestingly, some signatures were common to different types of cancer.
The other aspect of this study was to provide proof-of-concept that the proteomic analysis was a useful strategy to identify drivers of aggressive disease that could potentially be manipulated to control cancer growth.

“That’s exactly what we were able to do with this new, very powerful dataset,” said co-corresponding author Dr. Diana Monsivais, assistant professor of pathology and immunology at Baylor. “We focused on the uterine cancer data for which the computational analysis identified alterations in a number of proteins that were associated with aggressive cancer. We selected protein kinases, enzymes that would represent stronger candidates for therapeutics.”
Of hundreds of initial candidates, the researchers selected four kinases for functional studies in uterine cancer cell lines. They found that the kinases not only were expressed in the uterine cancer cells lines, but also that manipulating the expression of some of the kinases reduced the survival or the ability to migrate for some uterine cancer cells. Cell migration is a property of cancer cells that allows them to spread cancer to other tissues.
Teamwork
This study is the result of a productive collaboration between two Baylor centers, the Dan L Duncan Comprehensive Cancer Center and the Center for Drug Discovery.
“Chad conducted this wonderful analysis on new CPTAC datasets and was interested in validating it in a human cancer. He approached us about conducting functional studies to determine whether some proteins could translate to new targets for endometrial cancer,” Monsivais said.
Our experiments provided proof-of-concept that proteomics analysis is a useful strategy not only to better understand what drives cancer, but to identify new ways to control it or eliminate it,” Monsivais said.
In this study, we compared mRNA and protein signatures and, although in many cases they overlapped, about half the proteins in the proteomic signatures were not included in the corresponding mRNA signature, suggesting the need to include both mRNA and protein data in cancer studies,” Creighton said.
The researchers have integrated CPTAC data with the UALCAN data (An online portal for facilitating tumor subgroup gene expression and survival analyses), allowing users to query proteins of interest for associations with cancer grade or stage (http://ualcan.path.uab.edu/).
The study appears in the journal Oncogene.
Other contributors to this work include Yasmin Vasquez, Fengju Chen, Yiqun Zhang, Darshan S. Chandrashekar, John C. Faver, Ramya P. Masand, Michael E. Scheurer, Sooryanarayana Varambally and Martin M. Matzuk. The authors are affiliated with one or more of the following institutions: Baylor College of Medicine, University of Alabama at Birmingham, Texas Children’s Hospital and the University of Texas MD Anderson Cancer Center, Houston.
This work was supported by National Institutes of Health grants P30CA125123, P20CA221729, R00HD096057, a Core Facility Support Award from the Cancer Prevention Research Institute of Texas (RP160805) and a PDEP Award from the Burroughs Wellcome Fund. This research was conducted using data made available by the Clinical Proteomic Tumor Analysis Consortium, the Children’s Brain Tumor Tissue Consortium and the Cancer Genome Atlas Consortium.