Mystery solved, rotavirus VP3 is a unique capping machine
After eluding researchers for more than 30 years, the VP3 protein of rotavirus has finally revealed its unique structure and function to a team led by the labs of Dr. B. V. Venkataram Prasad and Dr. Mary K. Estes at Baylor College of Medicine.
The researchers knew that VP3 was necessary for capping messenger-RNA (mRNA), a process essential for the synthesis of viral proteins and for evading the host’s immune response, but while the structure and function of the other rotavirus proteins had progressively been unveiled, VP3 remained the last unsolved rotavirus mystery.
“Viruses cannot replicate on their own. They take over the machinery of the cells they infect to produce viral particles,” said co-corresponding author Prasad, professor and Alvin Romansky Chair of the Verna and Marrs McLean Department of Biochemistry and Molecular Biology and professor of molecular virology and microbiology at Baylor.

One of the strategies viruses use to highjack the cells they invade involves capping or preparing viral mRNA so that it mimics the capped mRNA of the host cells. Capping of mRNA is an essential step to engage the cellular machinery that synthesizes proteins. Imitating the cell’s capping enables the virus to take over the host’s machinery to produce viral proteins.
Several viruses use this strategy, but in different ways. In the case of rotavirus, it was known that the viral protein VP3 was involved in capping, but for more than three decades determining the structure of VP3 and understanding how it carried out mRNA capping had been a challenge.
Rotavirus VP3 protein is a unique capping machine

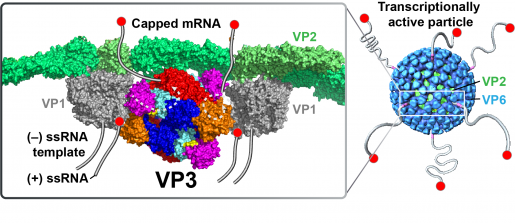
“Combining cryo-electron microscopy, X-ray crystallography and biochemical assays, we discovered that VP3 has all the enzymatic activities required to effectively cap rotavirus mRNA. We also found that, opposed to what has been seen in many viruses that have individual proteins for each enzymatic activity, rotavirus integrates all the enzymatic actitivities as modules into one protein, VP3,” said first author Dr. Dilip Kumar, postdoctoral fellow in the Prasad lab.

“What makes rotavirus different from most other viruses is that capping takes place inside the tight confines of the viral capsid, the protein shell of the virus, that is inside the cell. Once capping is complete, viral mRNA exits the capsid through channels and enters the cytoplasm of the cell where it engages the protein-making machinery to produce viral proteins,” said Estes, Cullen Foundation Endowed Chair and Distinguished Service Professor of molecular virology and microbiology at Baylor.
Solving this puzzle was very exciting,” Kumar said. “We were able to isolate VP3 and its modules and show that they self-assemble into a stable molecule capable of capping at least two mRNAs simultaneously, making the process efficient.”
Crucial to this work were the collaborations with Dr. Zhao Wang, assistant professor in the Verna and Marrs McLean Department of Biochemistry and Molecular Biology and co-director of the CryoEM Core at Baylor, and his postdoctoral fellow Dr. Xinzhe Yu as well as Dr. Sue Crawford, assistant professor in the molecular virology and microbiology for the enzymatic and functional assays, and Dr. Liya Hu, assistant professor in the Verna and Marrs McLean Department of Biochemistry and Molecular Biology, for the modeling of cryo-EM-data.
Next steps
Knowing the structure of VP3 and how it works offers the opportunity to design antiviral drugs to prevent or treat rotavirus infection. In addition, it makes it possible to further investigate the role VP3 plays in rotavirus replication and how it interferes with the host’s immune response.
“For nearly 30 years, my lab and Dr. Estes’s have collaborated to unravel the complex structure of rotavirus and better understand how it works,” Prasad said. “Determining the structure and function of VP3 is a major milestone that opens new doors for continuing our collaboration.”
Other contributors to this work include Ramakrishnan Anish, Soni Kaundal, Rodolfo Moreno and Joanita Jakana, all at Baylor, and Banumathi Sankaran at Lawrence Berkeley National Laboratory in California.
Find the report in the journal Science Advances.
This work was supported by National Institutes of Health MERIT grants AI36040 and R01AI080656, the Robert Welch Foundation (Q1279 and Q1967), National Institute of General Medical Sciences and the Howard Hughes Medical Institute. Further support was provided by the Office of Basic Energy Sciences of the U.S. Department of Energy under contract no. DE-AC02-05CH11231, the Advanced Technology Cores (ATC) CryoEM/ET core at Baylor College of Medicine and by the Protein and Monoclonal Antibody Production Shared Resource at Baylor College of Medicine with funding from NIH Cancer Center Support Grant P30 CA125123.