Better understanding of craniofacial birth defects opens new roads for regenerative medicine
Craniofacial birth defects, including cleft lip and palate, are among the most common human congenital malformations. These craniofacial anomalies occur because of defects in neural crest cells, whose role is to give rise to the complex craniofacial region by generating multiple cell types, including bone, cartilage and the peripheral nervous system.
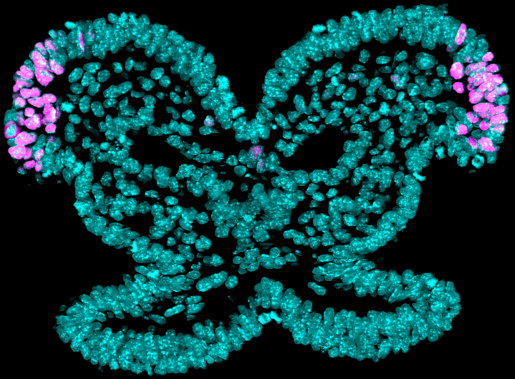
“Understanding neural crest cell development will allow us to better comprehend and treat those birth defects,” said first author of the work Rachel A. Keuls, graduate student in the lab of Dr. Ronald J. Parchem at Baylor College of Medicine. “In this study, we investigated what made it possible for neural crest cells to differentiate into a large variety of cell types during early development, as this could lead to strategies to generate healthy cells to repair craniofacial defects in human patients.”

“We discovered that neural crest cells have the ability to turn into many different cell types because they have access to multiple developmental programs,” Keuls said.
Using multiple approaches, the team uncovered that the use of many developmental programs results from chromatin in neural crest cells being more accessible than expected. Chromatin, a combination of genetic material and proteins, not only compacts the genome into the nucleus, but also is used by cells to control how the genome is interpreted to make different cell types.
“Bone cells emerge, for example, when chromatin opens giving access to genes that enable the cells to look like and conduct the functions of bone cells, while compact chromatin keeps other developmental programs leading to other cell types inaccessible,” Keuls said.
“The transient state in which chromatin is much more accessible enables neural crest cells to activate many genetic programs, which leads to the generation of multiple cell types required to develop the craniofacial region,” said Parchem, assistant professor of molecular and cellular biology and the Stem Cells and Regenerative Medicine Center at Baylor. He also is the corresponding author of the work.

The researchers then investigated what mediated the increased accessibility to chromatin in neural crest cells.
Previous evidence had shown that a class of small genetic material known as microRNAs can change cellular identity.
“We discovered that changes in chromatin accessibility in neural crest cells are regulated by the miR-302 microRNA family, which is highly expressed in cranial neural crest cells,” Keuls said.
Loss of miR-302 leads to reduced chromatin accessibility and a reduction in peripheral neuron differentiation. Mechanistically, the team found that miR-302 mediates the repression of multiple genes involved in chromatin condensation to promote accessibility required for cell differentiation.
Based on our findings, we hypothesize that disruption of the miR-302 regulatory axis may underlie some human craniofacial and neurological disorders,” Parchem said.
“From a basic science point of view, we would like to understand how this microRNA has this remarkable ability to alter developmental potential and chromatin accessibility. This is applicable to any field of stem cell biology.”
“We also see the possibility of translating our findings into clinical applications,” Keuls said.
The study has revealed that miR-302 is a powerful tool to modify stem cells to increase their ability to become a variety of cell types. We can potentially apply this ability to regenerate cells of the craniofacial region,” Keuls said.
Read all the details of this work in the Proceedings of the National Academy of Sciences.
Young Sun Oh and Ivanshi Patel, both at Baylor College of Medicine, also contributed to this work.
This project was supported by funding from the CPRIT Core Facility Support Award (CPRIT-RP180672), the NIH (CA125123 and RR024574) and by the Optical Imaging & Vital Microscopy Core, the Single-Cell Core, the Mouse Embryonic Stem Cell Core and the Genetically Engineered Mouse Core (P30 CA125123) at Baylor College of Medicine. Further support was provided by NIH grants (R01-HD099252, R01-HD098131, T32 ES027801-04, T32 HL092332), CPRIT Scholar in Cancer Research RR150106, Andrew McDonough B+ Foundation and V. Scholar in Cancer Research #V2017-017.