New method allows for a closer look at the effect of endocrine disrupting chemicals on gene expression
When something disrupts hormone binding to the receptor, such as environmental compounds or pollutants, called endocrine disrupting chemicals (EDCs), the results can be harmful and, in some cases, lead to certain forms of cancer. But exactly how certain compounds interfere with this process is not always easy to decipher.

Researchers at Baylor College of Medicine and Texas A&M University have further optimized a model cell line that was engineered to facilitate image analysis of multiple mechanistic steps of ER-stimulated gene transcription. The team developed new live cell quantitative analysis of ER dynamics using fluorescence recovery after photobleaching (FRAP) following exposures to EDCs.
“In past studies, we looked strictly at mobility – whether or not the receptor was moving when exposed to a hormone – using only a small number of known compounds since the assay was slow,” said Dr. Michael Mancini, professor of molecular and cellular biology at Baylor and an adjunct faculty member at the Texas A&M Health Science Center Institute for Biosciences and Technology (IBT).

As reported earlier, the approach was able to link changes in ER mobility to known activating or inactivating hormones.
Faster and more accurate method allows for deeper analysis
“In the current work, we now see much more detail—for example, how much receptor is in a cell and whether it is engaged at the visible multicopy reporter gene locus—and now this is fast enough to facilitate testing a larger group of known hormones and EDCs,” Mancini said.

Using a panel of 45 compounds known to target ER, provided by the Environmental Protection Agency, and supported in part by an NIEHS-supported, Texas A&M Superfund Research Center (P42ES027704), researchers began using a new, more sensitive version of the cell model along with FRAP to take a closer look at how receptor mobility relates to transcription.
“We were able to see, for example, which compounds result in the receptors targeting the transcription locus and whether they repressed or activated transcription from that locus,” said Dr. Michael Bolt, assistant research scientist with Texas A&M. “One interesting observation was that while some compounds activated ER, the amount of activation would plateau. Despite the same level of transcriptional activation, some compounds caused the receptor to move slowly and others quickly. We’re unclear why at present, or where in the cascade of steps this could have an effect.”
While these observations are important to understanding how environmental factors are affecting ER, the goal of this research was to create and fine tune a process to effectively test more compounds.
This brought together the use of the new cell line with proven high content analysis approaches, and also combined with FRAP using a newly-developed algorithm to greatly speed-up the process and analysis of larger numbers of cells.
“Most FRAP approaches test only a limited number of cells in a microscope field at a time. So, if you wanted to test a lot of cells for better statistics, that would require many different imaging runs,” Mancini said. “Now, with informatics support from our CPRIT core resource grant that employs both Baylor and IBT lab members, we have a new way to compare a live image series, using image math to find and analyze multiple bleached regions at a time, making the process much more efficient. Drs. Bolt and Pankaj Singh, who developed the new analysis pipeline, have nicknamed this process as ‘FRAPnalysis.’”
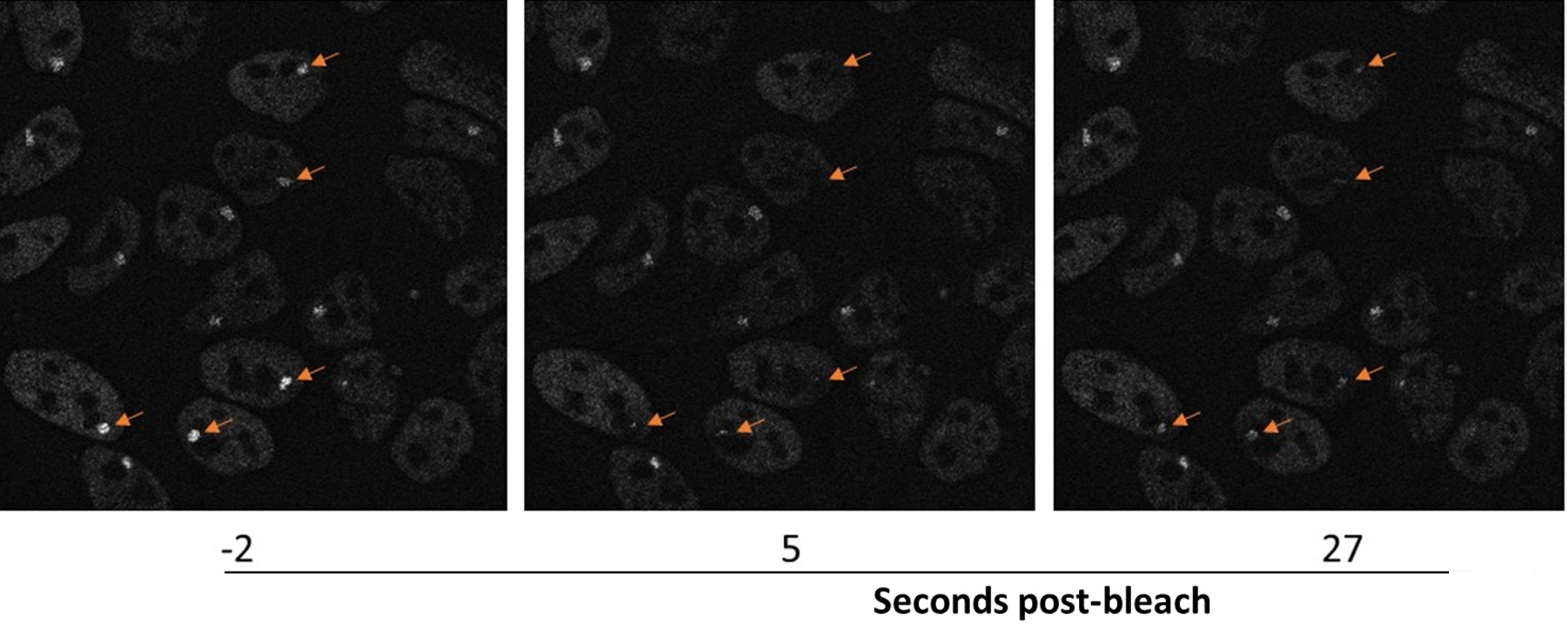
Mancini said these robust assay developments are opening-up the field of environmental testing due to increased speed and accuracy.
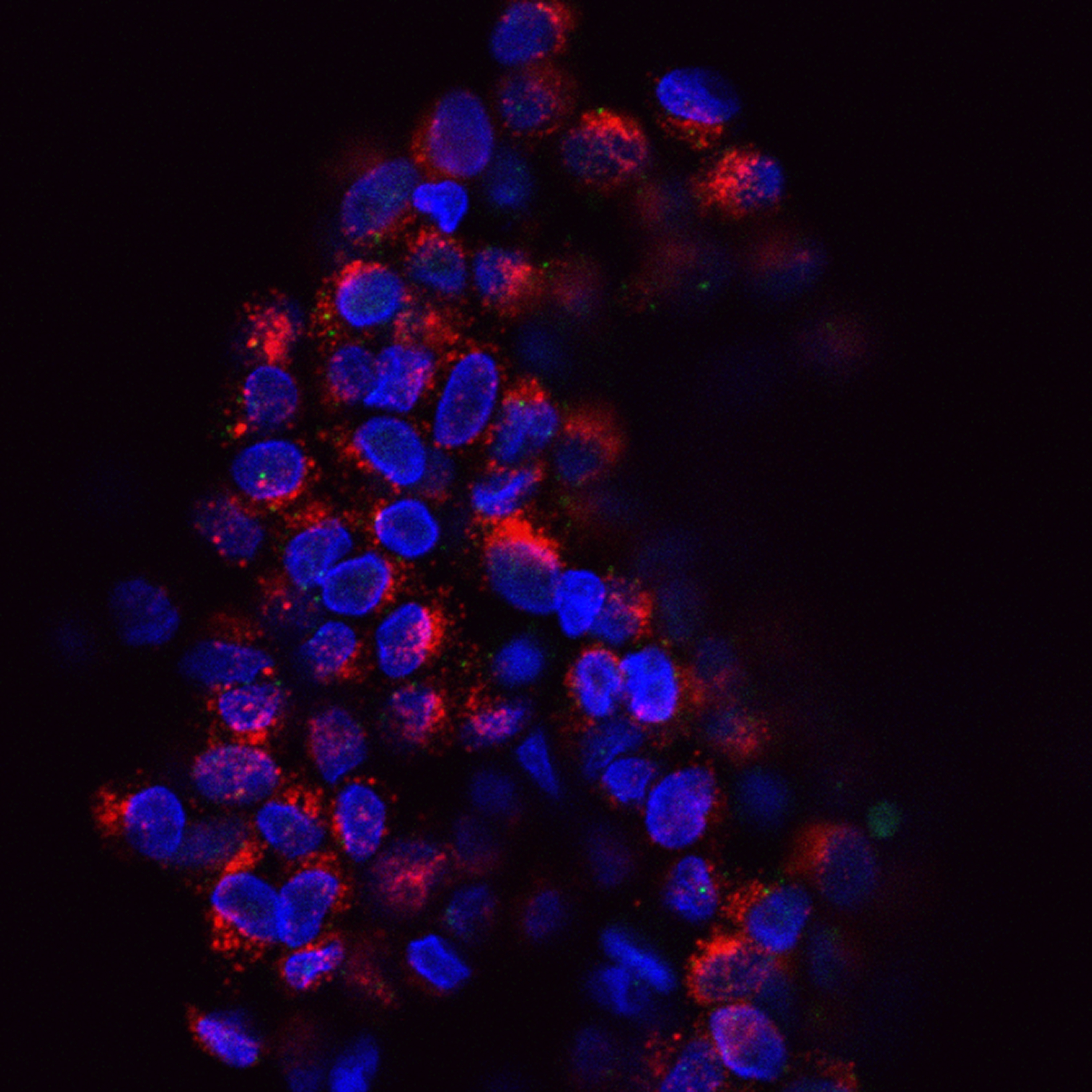
“We have created panel of cell lines that have green florescent protein-tagged estrogen (and other) receptors and a visible transcription locus in nuclei. Without ligands, the nuclear ER signal looks like a smooth sea of glass. However, if you add a compound, we literally can see a spot form in minutes, telling you something is going on with the receptor at the transcription locus. Whether a compound changes the size of the reporter gene locus is linked to how it affects transcription. Overall, we continue to see and quantify the process in more detail than before,” Mancini said.
Bolt said they can get certain results and information in less than an hour and the amount of data that is being processed is much greater than before. “What we are seeing now is that the dynamics of ERα bound to different classes of compounds are much more complicated than we thought,” he said.
The findings are described in the current edition of iScience.
Others who took part in this study include: P. Singh, C.E. Obkirchner and R.T. Powell, with Texas A&M, and A.T. Szafran, M.G. Mancini and F. Stossi with Baylor.
The work was partially supported by NIEHS (P42ES027704), the CPRIT-funded Gulf Coast Consortia (GCC) Center for Advanced Microscopy and Image Informatics (RP170719), the CPRIT-funded Combinatorial Drug Discovery Program (RP200668) and the GCC Center for Precision Environmental Health (P30ES030285).