How estrogen regulates gene expression
Estrogens are a group of hormones that are essential for normal female sexual development and for the healthy functioning of the reproductive system. They also are involved in certain conditions, such as breast cancer. Estrogen also plays a role in male sexual function. These hormones carry out their functions by turning genes on and off via a multi-step process. After estrogen binds to its receptor in the cell nucleus, different coactivators bind to the complex in a sequential manner.
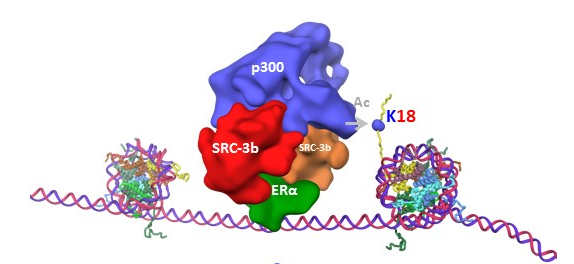

“Experimental evidence suggests that different estrogen-receptor coactivators communicate and cooperate with each other to regulate gene expression,” said corresponding author Dr. Bert O’Malley, chair and professor of molecular and cellular biology and Thomas C. Thompson Chair in Cell Biology at Baylor College of Medicine. “However, how this communication takes place and how it guides the sequence of events that regulate gene expression was not clear.”
The study shows that the sequential recruitment of coactivators is not simply adding molecules to the complex, it results in dynamic specific structural and functional changes that are necessary for effective regulation of gene expression.
O’Malley, Dr. Wah Chiu, Distinguished Service Professor and Alvin Romansky Professor of Biochemistry and Molecular Biology at Baylor during the development of this project, and their colleagues combined cryo-electron microscopy structure analysis and biochemical techniques and showed how the recruitment of a specific coactivator – CARM1 – into the complex guides the subsequent steps leading to gene activation.

“For the estrogen receptor complex to be able to regulate gene expression, the coactivator CARM1 needs to be added after other coactivators have been incorporated into the complex,” said first author Dr. Ping Yi, assistant professor of molecular and cellular biology at Baylor.
We discovered that when CARM1 is added, it changes the complex both chemically and structurally, and these changes guide subsequent steps that lead to gene activation.”
(See video of molecular interactions below. Credit: O’Malley lab and Chiu lab).
“We now have a better understanding of how this molecular machine works and of what role each one of the components plays. We are better prepared to understand what might have gone wrong when the machine fails,” O’Malley said.
Find all the details about this research in the journal Molecular Cell.
###
Other contributors to this work include Zhao Wang, Qin Feng, Chao-Kai Chou, Grigore D. Pintilie, Hong Shen, Charles E. Foulds, Guizhen Fan, Irina Serysheva, Steven J. Ludtke, Michael F. Schmid, Mien-Chie Hung and Wah Chiu. The researchers are affiliated with one or more of the following institutions, researchers at Baylor College of Medicine, the University of Texas MD Anderson Cancer Center and the University of Texas Health Science Center at Houston.
Support for this study was provided by the Komen Foundation (5PG12221410), the Department of Defense (R038318-I and W81XWH-15-1-0536); National institutes of Health grants (HD8818, NIDDK59820, P41GM103832 and R01GM079429); CNIHR, R21AI122418 and R01GMGM072804; CPRIT grants (RP150648 and DP150052); and a National Cancer Institute Cancer Center Support grant (P30CA125123) to the BCM Monoclonal Antibody/recombinant Protein Expression Core Facility.