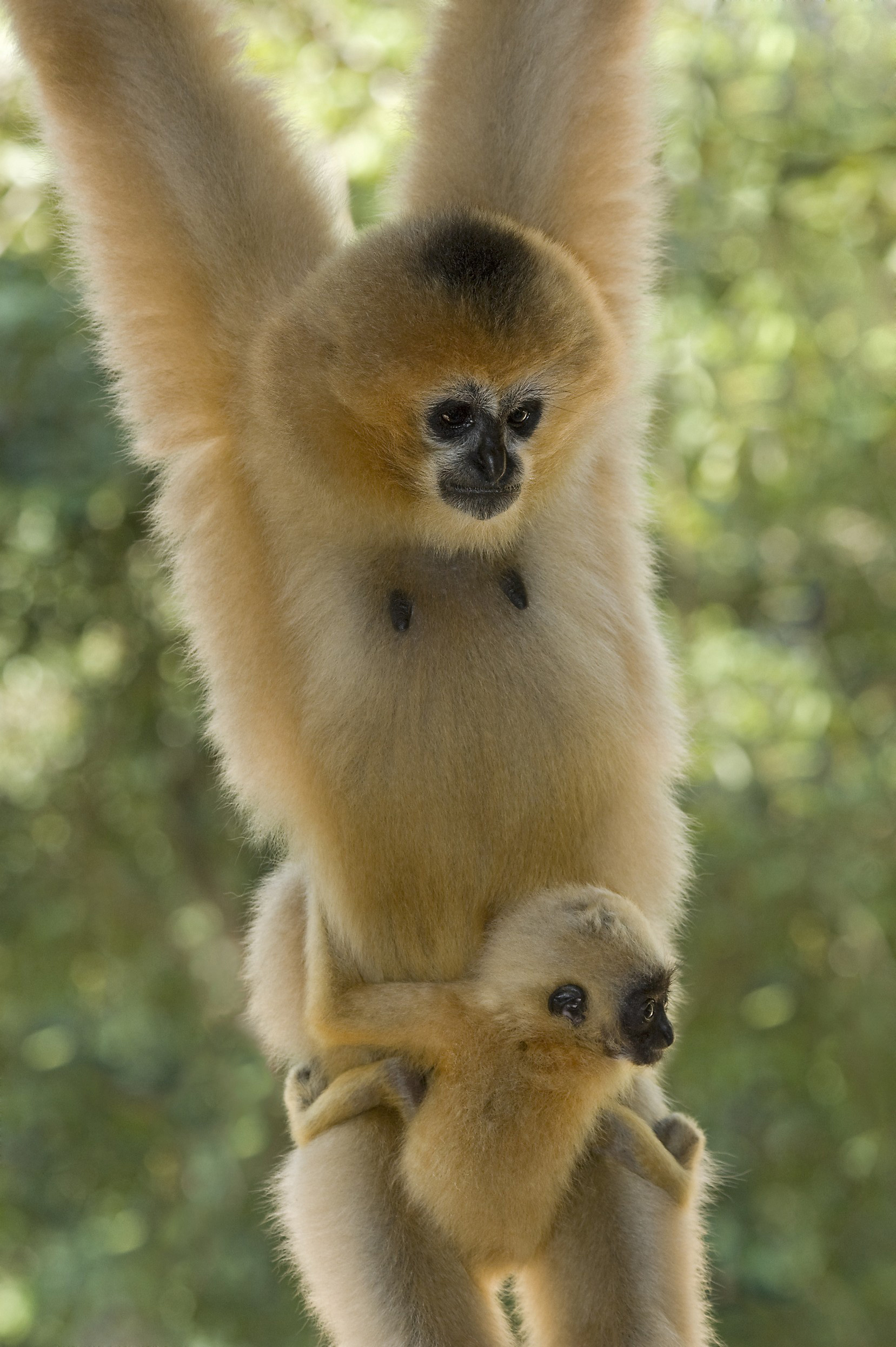
Genome sequence of nimble gibbon
The gibbon nimbly maneuvers through the branches of his or her environment, but until an international consortium of experts that included those at the Baylor College of Medicine Human Genome Sequencing Center sequenced and analyzed the ape’s genome, scientists were unclear on the effects of its unusually rapid rate of chromosome rearrangement.
In a report in the journal Nature , researchers led by those from Baylor College of Medicine, Oregon Health & Science University and Washington University in St. Louis, Mo., describe the rapid rate of chromosomal rearrangements in this ape, a fact that has implications for human genetic diseases and cancer. While such rearrangements of the chromosomes – the packaging that encases the genetic information stored in the DNA sequence – do not appear to cause problems for gibbons, it does for humans.
“Everything we learn about the genome sequence of this particular primate and others analyzed in the recent past helps us to understand human biology in a more detailed and complete way,” said Dr. Jeffrey Rogers, associate professor in the Human Genome Sequencing Center at Baylor and a lead author on the report.

“The gibbon sequence represents a branch of the primate evolutionary tree that spans the gap between the Old World Monkeys and great apes and has not yet been studied in this way. The new genome sequence provides important insight into their unique and rapid chromosomal rearrangements.”
For years, experts have known that gibbon chromosomes evolve quickly and have many breaks and rearrangements, but up until now there has been no explanation why, Rogers said. The genome sequence helps to explain the genetic mechanism unique to gibbons that results in these large scale rearrangements.
The sequencing was led by Dr. Kim Worley, professor in the Human Genome Sequencing Center, and Rogers, both of Baylor and Drs. Wesley Warren and Richard Wilson of Washington University.

The analysis was led by Dr. Lucia Carbone, an assistant professor of behavioral neuroscience in the OHSU School of Medicine and an assistant scientist in the Division of Neuroscience at OHSU’s Oregon National Primate Research Center. Carbone is an expert in the study of gibbons and the lead and corresponding author on the report.
“We do this work to learn as much as we can about gibbons, which are some of the rarest species on the planet,” said Carbone. “But we also do this work to better understand our own evolution and get some clues on the origin of human diseases.”

Chromosomes play an essential role in the packaging of DNA, Worley said. “There are 3 billion base pairs of DNA in every cell and it is packaged into 23 pairs of chromosomes,” said Worley, also a lead author on the report.
When there are rearrangements in chromosomes, the genes and gene regulation are often disrupted or broken, said Worley.
“Cancer is clearly the biggest medical example of the impact of chromosome rearrangements. The gibbon sequence gives us more insight into this process,” said Worley. “There are also a number of other genetic diseases that result from these events.”
Rearrangements
The number of chromosomal rearrangements in the gibbons is remarkable, Rogers said. “It is like the genome just exploded and then was put back together,” he said. “Up until recently, it has been impossible to determine how one human chromosome could be aligned to any gibbon chromosome because there are so many rearrangements.”
Repeat element
The sequencing project revealed a novel and unique genetic repeat element (segments of DNA that occur in multiple copies throughout the genome) that is inserted into genes associated with the maintenance of chromatin structure.
Repeat elements have the capability to disrupt a gene and change their biological function, Worley said. In the gibbons, the team identified the LAVA element – a novel repeat element that emerged exclusively in gibbons and preferentially hits genes involved in chromosome segregation (an essential step in cell division, where chromosomes pair off with their similar homologous chromosome.)
“This explains why the gibbons have undergone so much change,” said Rogers. “Similar disruptions cause disease, which is why everything we learn from this helps us better understand human biology and chromosomal structures.”
Collaborating institutions include Nabsys Inc., in Providence, RI; University of Arizona in Tucson; University of Pompeu Fabra/CSIC, ICREA at Institut de Biologia Evolutiva, Barcelona, Spain; University of Washington,Seattle, WA; Howard Hughes Medical Institute; European Molecular Biology Laboratory- European Bioinformatics Institute, Wellcome Trust Genome Campus in Hinxton, United Kingdom; Leibniz Institute for Primate Research, Gene Bank of Primates, German Primate Center, Göttingen, Germany; Wellcome Trust Sanger Institute, Wellcome Trust Genome Campus in Hinxton, United Kingdom; University of Bari, Dipartimento di Biologia, Bari, Italy; Louisiana State University, Department of Biological Sciences, Baton Rouge, LA; University of Paul Sabatier, Toulouse, France; The Johns Hopkins University School of Medicine, Department of Molecular Biology and Genetics, Baltimore, MD; University of Utah, Salt Lake City; Texas A&M University, Department of Ecosystem Science and Management, College Station; Babes-Bolyai-University, Institute for Interdisciplinary Research in Bio-Nano-Sciences, Molecular Biology Center, Cluj-Napoca, Romania; Children’s Hospital Oakland Research Institute, BACPAC Resources, Oakland, Calif; University of Colorado School of Medicine, Department of Biochemistry and Molecular Genetics, Aurora, CO; Delbrück Center for Molecular Medicine, Berlin, Germany; Centro Nacional de Análisis Genómico (CNAG), Barcelona, Spain; Indiana University School of Informatics and Computing, Bloomington, IN; Washington University School of Medicine, The Genome Institute, St. Louis, MO; Institute for system biology, Seattle; The Pennsylvania State University, Department of Anthropology, University Park, PA; University of Pittsburgh School of Medicine, Department of Developmental Biology-Department of Computational and Systems Biology, Pittsburg, PA; Harvard Medical School, Department of Genetics, Boston, MA; University of Cambridge, Cancer Research UK-Cambridge Institute, Cambridge, United Kingdom; University of San Francisco, Gladstone Institutes, Institute for Human Genetics – Department of Epidemiology & Biostatistics, San Francisco, Calif; Paul Ehrlich Institute, Division of Medical Biotechnology, Lagen, Germany; Gibbon Conservation Center, Santa Clarita, CA; Louisiana State University, School of Electrical Engineering and Computer Science, Baton Rouge, LA; University of California San Francisco, Institute for Human Genetics, San Francisco, CA; Stony Brook University, Department of Ecology and Evolution, Stony Brook, NY.
Funding for this work was provided by the National Human Genome Research Institute (U54 HG003273; U54 HG003079; National Institutes of Health ( P30 AA019355 and NIH/NCRR P51 RR000163; R01_HG005226; National Library of Medicine Biomedical Informatics Research Training Program (R01 GM59290, U41 HG007497-01, R01 MH081203, HG002385; National Science Foundation (CNS-1126739); European Research Council (260372): Ministerio de Ciencia e Innovacion (BFU2011-28549): Ministry of National Education (PN-II-ID-PCE-2012-4-0090: EMBO Young Investigator Award; University of Cambridge; Cancer Research UK; Commonwealth Scholarship Commission.
— Glenna Picton